Can you sequence an antibody without hybridoma cells?expand_more
Yes. If purified antibody protein is available, LC-MS/MS-based de novo antibody sequencing can be used without hybridoma cells, DNA, or RNA. This is suitable for unknown antibodies, commercial antibodies, and lost hybridoma projects.
Can you sequence a commercial antibody?expand_more
Yes. Commercial antibodies can often be analyzed by LC-MS/MS-based antibody sequencing if sufficient material and acceptable sample quality are available. Providing the datasheet, clone name, host species, concentration, formulation, and antibody type helps with project evaluation.
What is de novo antibody sequencing?expand_more
De novo antibody sequencing reconstructs antibody amino acid sequences without relying on a complete reference DNA or protein sequence. In LC-MS/MS-based workflows, antibody proteins are digested into peptides, analyzed by tandem mass spectrometry, and assembled into heavy and light chain sequences.
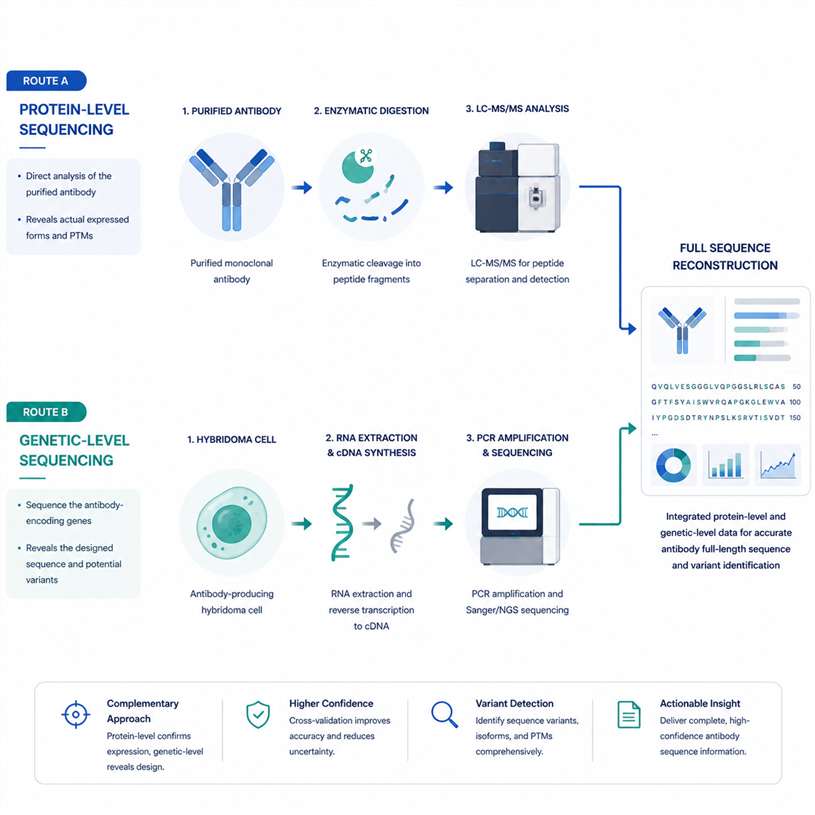
What is the difference between LC-MS/MS-based and PCR-based antibody sequencing?expand_more
LC-MS/MS-based antibody sequencing starts from antibody protein and does not require genetic material. PCR-based antibody sequencing starts from hybridoma cells, B cells, RNA, or other genetic material. LC-MS/MS is preferred when only purified antibody is available, while PCR-based sequencing is preferred when viable antibody-producing cells are available.
How do Sanger sequencing and NGS fit into antibody sequencing?expand_more
Sanger sequencing can confirm amplified or cloned VH/VL sequences from defined antibody sources. NGS-supported antibody sequencing can support B-cell repertoire, antibody library, and discovery projects when genetic material is available.
Can you identify CDR regions?expand_more
Yes. CDR identification and annotation can be included when sufficient sequence information is obtained from heavy and light chain variable regions. CDR analysis is especially useful for antibody engineering, humanization, and antigen-binding studies.
Can you sequence both heavy and light chains?expand_more
Yes. Heavy and light chain sequence analysis can be performed using LC-MS/MS-based or genetic workflows. Chain-specific interpretation may require chain separation, overlapping peptide evidence, or genetic material depending on the project.
What sample type gives the best result?expand_more
Purified monoclonal antibody is generally the preferred sample type for LC-MS/MS-based de novo antibody sequencing. For PCR-based, Sanger, or NGS-supported sequencing, high-quality hybridoma cells, B cells, RNA, amplicons, or library material may be preferred.
Can antibody sequencing identify PTMs or modifications?expand_more
LC-MS/MS-based workflows can support protein-level modification analysis when included in the project scope. However, modification detection depends on sample quality, modification type, peptide coverage, and analytical strategy.
Can you distinguish leucine and isoleucine?expand_more
Leucine and isoleucine have the same mass, so they can be difficult to distinguish by standard MS/MS. When residue-level differentiation is critical, additional strategies or orthogonal evidence may be required. Ambiguities should be clearly reported in the final results.
What causes incomplete antibody sequence coverage?expand_more
Incomplete coverage can result from low sample amount, poor purity, difficult peptide behavior, incompatible buffer components, modifications, glycosylation, conjugation, or regions that do not generate suitable peptides under a given digestion condition.
What will be included in the final report?expand_more
The final report may include heavy and light chain sequences, VH/VL annotation, CDR annotation, peptide evidence, sequence coverage maps, confidence assessment, ambiguous residue notes, PTM information if applicable, and expert interpretation.
Creative Proteomics provides antibody sequencing services for research use only. These services are not intended for clinical diagnosis, treatment, or direct therapeutic decision-making.