- Home
-
Services
-
Protein Sequencing
- Mass Spectrometry Based Protein Sequencing
- Edman Based Protein Sequencing
- Nanopore Protein Sequencing
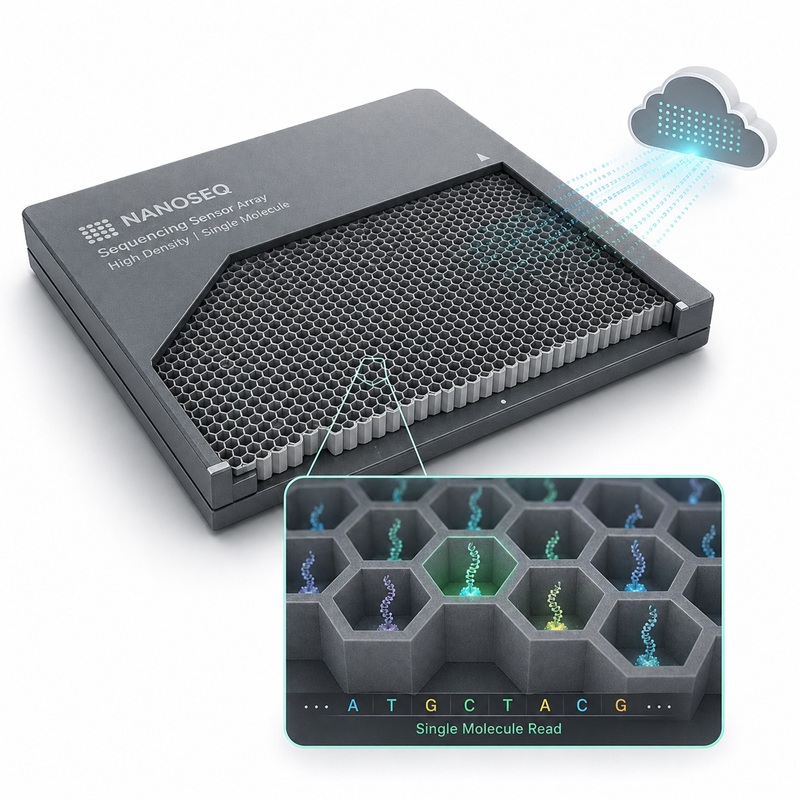
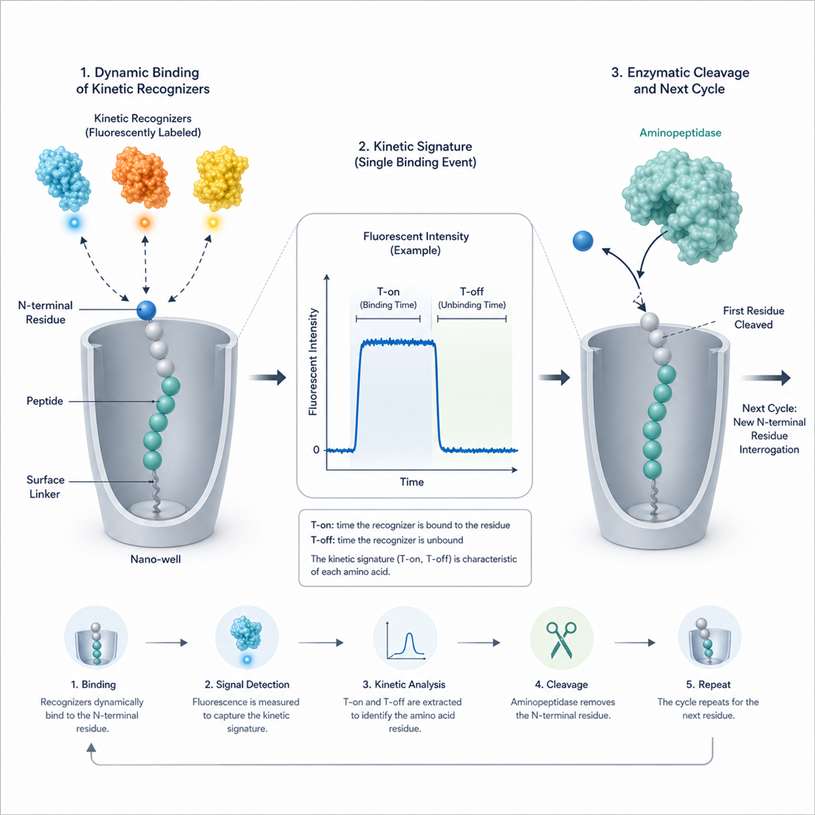
- Single Molecule Fluorescence Sequencing
-
Antibody Sequencing
- Mass Spectrometry Based Antibody Sequencing
- PCR Based Antibody Sequencing
- PhIP-Seq Antibody Analysis Service
- Peptide Sequencing
-
Protein Sequencing
-
Applications
-
Proteomics Analysis Services
- Protein Identification Services
-
Protein Post-Translational Modification Analysis Services
- Phosphorylated Protein Analysis Service
- Multi-Channel Phosphorylated Protein Analysis Service
- Acetylated Protein Analysis Service
- Ubiquitinated Protein Analysis Service
- Glycosylation Protein Analysis Service
- Histone Modification Analysis Service
- Disulfide Bond Analysis Service
- Methylation Protein Analysis Service
- Acylation Quantitative Protein Analysis Service
- Protein Sumoylation Identification Service
- Top-Down Based PTMs Characterization Services
-
Glycoproteomics Analysis Services
- Glycoprotein Analysis Services
- N-Glycan Analysis Service
- O-Glycan Analysis Service
- N-Glycosylation Site Analysis Service
- O-Glycosylation Site Analysis Service
- N-Glycan Modification and Modification Site Analysis Service
- O-Glycan Modification and Modification Site Analysis Service
- Glycopeptide Analysis Service
- Glycosphingolipid Analysis Service
- Top-Down Based Glycoprotein Analysis Service
- Peptidomic Analysis Services
- Biopharmaceutical Characterization Services
- Bioinformatics Services
-
Proteomics Analysis Services
- Company
- Resource
- Inquiry