What NGPro™ Includes: A Proteomics Platform Built for Discovery, Quantification, and Validation
Relying on multiple fragmented CROs for different phases of your research often leads to incompatible datasets, differing extraction efficiencies, and lost momentum. The NGPro™ platform solves this by offering an end-to-end translational pipeline. We ensure that candidate biomarkers identified in the discovery phase are smoothly and reliably transitioned into the validation phase using standardized quality control, matched hardware, and a unified bioinformatics logic.
Choose the Right Proteomics Route for Your Study
Selecting the appropriate mass spectrometry technology can be complex. To ensure you get the most actionable data, we have structured our platform around your specific project stage and sample limitations. Choose the route that best describes your current research challenge:

Route 1: Exploring Unknown Biology (Broad Discovery)
Mapping global protein expression profiles without prior knowledge or established target lists.
- Best for: Early-stage basic research, cell line profiling, drug mechanism exploration.
- Typical Samples: Cell cultures, generic tissue lysates.
- Key Output: Broadest possible protein ID list (up to 7,000+ proteins).
- Where to go next: Explore our unbiased discovery proteomics services.

Route 2: Cohort-Scale Quantitative Continuity
Achieving robust, gap-free statistical power across dozens or hundreds of independent clinical samples without batch effects.
- Best for: Large-scale biomarker screening, clinical cohort phenotyping, multi-center studies.
- Typical Samples: Plasma, serum, urine, large tissue cohorts.
- Key Output: Highly consistent quantitative matrices with minimal missing values.
- Where to go next: Learn about our DIA quantitative proteomics service, 4D-DIA proteomics platform, or SWATH-MS digital archiving.
Route 3: Ultra-Low Input & Complex Matrices
Profiling precious, finite samples or highly complex backgrounds where low-abundance signals are easily masked.
- Best for: Micro-scale biopsies, spatial proteomics, rare immune cell subtyping.
- Typical Samples: FACS-sorted cells, Laser Capture Microdissection (LCM) tissues, FFPE curls.
- Key Output: Extreme identification depth extracted from nanograms of input via 4D ion mobility separation.
- Where to go next: Discover our low-input 4D label-free proteomics services.

Route 4: Targeted Clinical Verification
Performing absolute, assay-grade quantification for a predefined target list (e.g., 10 to 150 proteins).
- Best for: Biomarker validation, clinical assay development, confirming discovery hits, replacing ELISA/Western Blots.
- Typical Samples: Clinical biofluids, drug-treated tissues.
- Key Output: Absolute molar quantification (e.g., fmol/μL) with extremely high specificity.
- Where to go next: Review our custom targeted proteomics services, including PRM quantitative proteomics, 4D-PRM target proteomics, and SRM-MRM quantitative workflows.
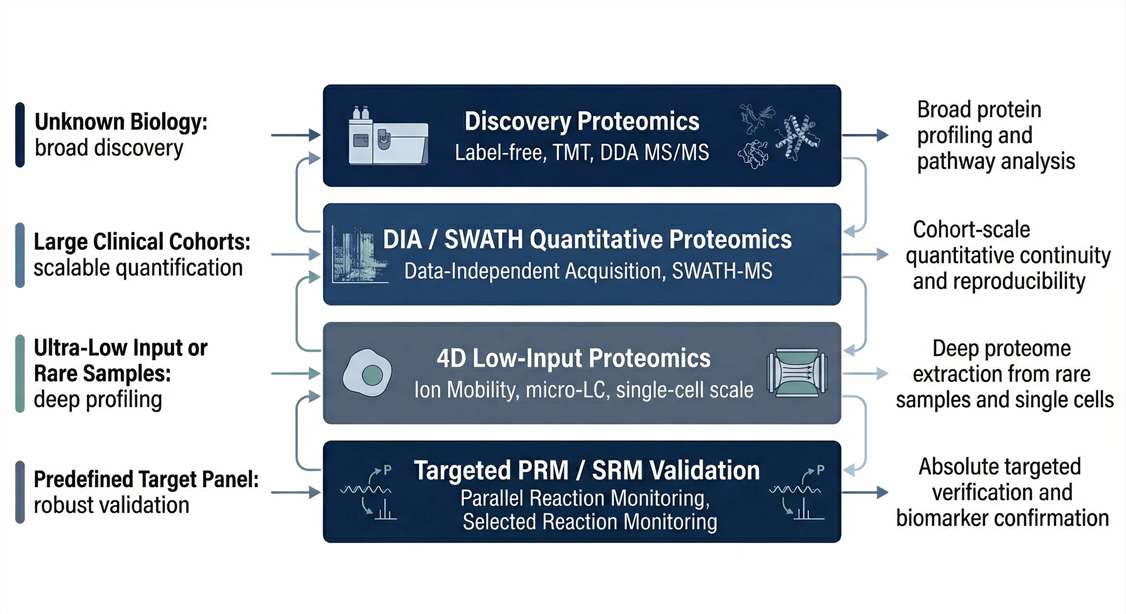
Technology Selection Guide: Discovery vs. DIA vs. 4D vs. Targeted Proteomics
Once your route is determined, mapping it to the exact mass spectrometry technology is critical for project success.
| Dimension | Traditional DDA (Broad Discovery) | DIA / SWATH (Cohort Screening) | 4D Proteomics (Low-Input/Depth) | Targeted PRM / SRM (Validation) |
|---|---|---|---|---|
| Primary Objective | Maximum raw IDs in single/small sets | Missing-value elimination across cohorts | Deep extraction from trace amounts | Absolute quantification of specific targets |
| Throughput (Proteins) | Up to 7,000+ per run | Up to 7,000 - 9,000+ per run | 5,000+ from nanogram inputs | Highly specific panel (10 - 150) |
| Sensitivity Focus | Standard MS1 intensity | Continuous MS2 recording | Physical CCS isomer separation | Attomole-level limit of detection |
| Cohort Scalability | Low (Stochastic dropouts) | Infinite (Highly reproducible) | Excellent | High (Multiplexed targeting) |
| When NOT to choose | When you need reproducible quantification across large cohorts. | When your sample amount is extremely limited (<1 μg). | When absolute molar quantification of known targets is required. | For unbiased, hypothesis-free discovery of new markers. |
Instrumentation and Technologies for Proteomics Discovery and Validation
The NGPro™ platform utilizes a meticulously integrated mass spectrometry infrastructure. We do not force projects onto a single instrument. Instead, we deploy a multi-vendor elite fleet to leverage the distinct strengths of each architecture:
Bruker timsTOF Pro Series
Deployed for 4D proteomics, ultra-low input samples, and complex 4D protein post-translational modifications (PTMs) proteomics where Trapped Ion Mobility Spectrometry (TIMS) is required to physically separate co-eluting background noise.

Thermo Scientific Orbitrap Series
Deployed for high-resolution discovery, deep global profiling, and applications requiring extreme mass accuracy and wide dynamic range.

SCIEX TripleTOF & Triple Quad Series
Deployed for highly stable, large-scale SWATH-MS acquisition and robust, multiplexed targeted SRM/MRM quantification.
How Our Proteomics Workflow Moves Projects from Study Design to Biological Interpretation
Our standardized operating procedures ensure high reproducibility and traceability across all NGPro™ services.
Tailoring the analytical route (Discovery, DIA, 4D, or Targeted) and statistical power to your biological question.
Performing optimized lysis and digestion. Strict QC checkpoints, including indexed Retention Time (iRT) spike-ins, assess peptide yield and purity before mass spectrometry.
High-throughput LC-MS/MS acquisition with continuous instrument tuning and batch harmonization controls to prevent run-to-run drift over multi-month studies.
Utilizing spectral libraries or library-free algorithms with rigorous False Discovery Rate (FDR) control (<1%) to ensure unassailable data matrices.
Delivering customized biostatistical modeling, pathway enrichment, and publication-ready visualizations.
Sample Preparation Capabilities and Requirements
A high-end mass spectrometer cannot rescue poor sample extraction. We tie our preparation protocols directly to matrix difficulty, offering specialized FFPE quantitative proteomics solutions, automated high-abundance protein depletion for blood, plasma, and serum proteomics solutions, and low-loss micro-extraction for rare cells.
Specific input recommendations vary based on the selected analytical strategy:
| Sample Matrix | Discovery / DIA Input | Ultra-Low Input (4D) | Targeted Input (PRM/MRM) | Preservation |
|---|---|---|---|---|
| Plasma / Serum | 200 μL | Not typically recommended | 50 - 100 μL | Flash-frozen, avoid freeze-thaw |
| Tissue (Fresh/Frozen) | 10 - 20 mg | < 1 mg (Biopsy cores) | 5 - 10 mg | Snap-frozen, dry ice |
| FFPE Slices | 5 - 10 curls (10μm) | 1 - 2 curls | 2 - 5 curls | Room temp / slide mailer |
| Cells | 1×10&sup7; cells | 1,000 - 10,000 cells | 5×10&sup6; cells | Washed pellet, snap-frozen |
Bioinformatics Outputs That Turn Protein Lists into Decisions
We provide specialized proteomics bioinformatics analysis services designed to translate raw mass spectrometry spectra into actionable insights, focusing on the critical handoff from broad discovery to targeted validation.
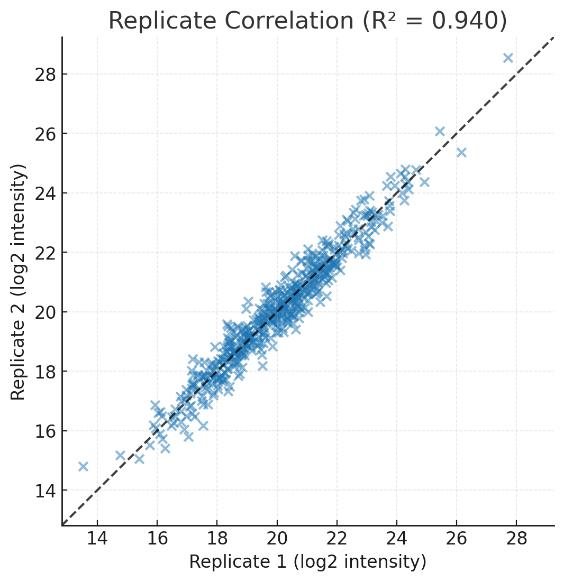
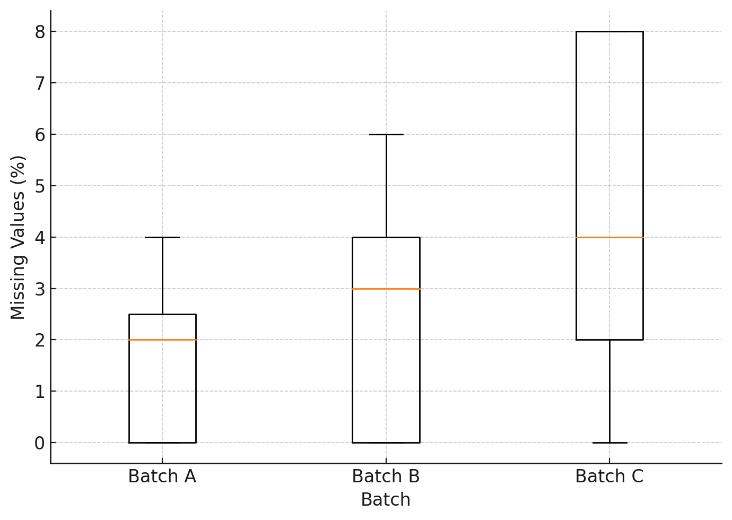
Cohort Stability: PCA and CV density plots demonstrating tight clustering of biological replicates and the elimination of batch effects across large-scale runs.
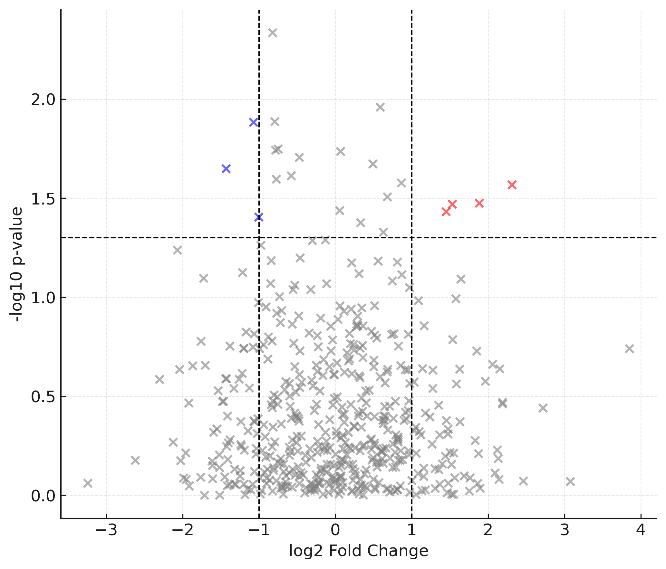
Biomarker Screening: High-resolution Volcano plots and hierarchical clustering heatmaps clearly delineating statistically significant up- and down-regulated protein panels.
Machine Learning Prioritization: ROC curve matrices and LASSO regression plots utilized to evaluate the predictive accuracy of selected biomarker candidates.
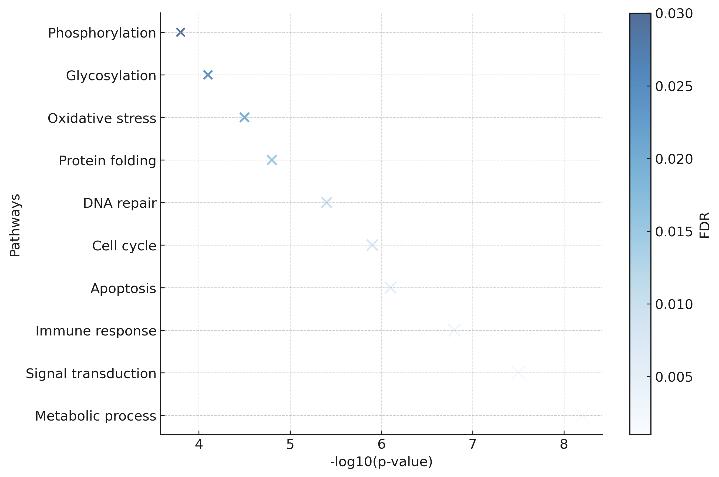
Functional Enrichment: KEGG, GO, and Reactome pathway mapping to explain the biological mechanisms driving the observed proteomic shifts.
Raw Data Files
Native instrument files for independent archival and verification.
Processed Quantitative Matrices
Clean, missing-value-filtered tables ready for secondary analysis.
Comprehensive QC Report
Detailed documentation of extraction efficiency, ID rates, and instrument parameters.
Publication-Ready Figures
High-resolution graphs and statistical interpretations ready for manuscript integration.
Applications Across Biomarker Discovery, Translational Research, and Drug Development
The NGPro™ platform is engineered to resolve high-value, complex project scenarios where standard proteomics falls short, bridging the gap between exploratory science and clinical utility:
- Biomarker Discovery to Validation: Executing the complete pipeline from screening tissue/tumor microenvironments to validating circulating markers in biofluids. Explore our comprehensive biomarker proteomics solutions, including targeted biofluid biomarker discovery solutions and deep tissue biomarker discovery solutions.
- Translational Cohort Studies: Managing massive clinical datasets with robust quantitative continuity for disease phenotyping. Learn more about our clinical proteomics research solutions.
- Drug Mechanism and Target Deconvolution: Identifying protein binding partners, elucidating mechanisms of action, and assessing off-target toxicities of novel compounds. See our approaches for drug target identification solutions and proteomics solutions of drug mechanism of action.
- Low-Input and Rare-Sample Proteomics: Digitizing the proteomes of precious, volume-limited samples. Discover our workflows for exosomes proteomics research solutions and spermatozoa proteome research solutions.
- Immunology and Microbiome Profiling: Mapping the immune landscape via immunopeptidome profiling services or understanding complex microbial communities through 4D-LFQ metaproteomics services.
Frequently Asked Questions About Choosing the Right Proteomics Workflow
Case Study: Simultaneous Discovery and Targeted Clinical Proteotyping with Hybrid PRM/DIA
Journal: Clinical Proteomics · Published: 2024
Study Scope
Clinical proteomics projects often face a practical bottleneck when discovery-stage screening and targeted validation are performed as two separate workflows. This can create inconsistencies in quantitative logic, increase sample consumption, and slow the transition from broad candidate identification to focused biomarker verification.
In this study, researchers developed a hybrid PRM/DIA workflow that integrates targeted peptide monitoring with discovery-driven proteome profiling in a single LC-MS acquisition strategy.
- The workflow was first evaluated using 185 proteotypic peptides derived from 64 annotated protein groups.
- For targeted triggering, the authors incorporated 30 AQUA peptides representing 28 biomarker candidates.
- The method was then applied to biobanked melanoma samples in a clinically relevant proteotyping setting.
Quantitative Performance and Workflow Features
The hybrid PRM/DIA workflow demonstrated that targeted monitoring could be added without sacrificing broad proteome coverage.
- The targeted PRM layer improved the reproducibility and sensitivity of low-abundance endogenous peptide detection.
- Up to 179 MSxPRM scans were introduced without compromising overall DIA performance.
- In the melanoma samples, the DIA component still detected approximately 6,500 protein groups, preserving broad proteome context while enabling focused monitoring of selected biomarker candidates.
- Proteins such as UFO, CDK4, NF1, and PMEL were consistently monitored through the integrated targeted layer.
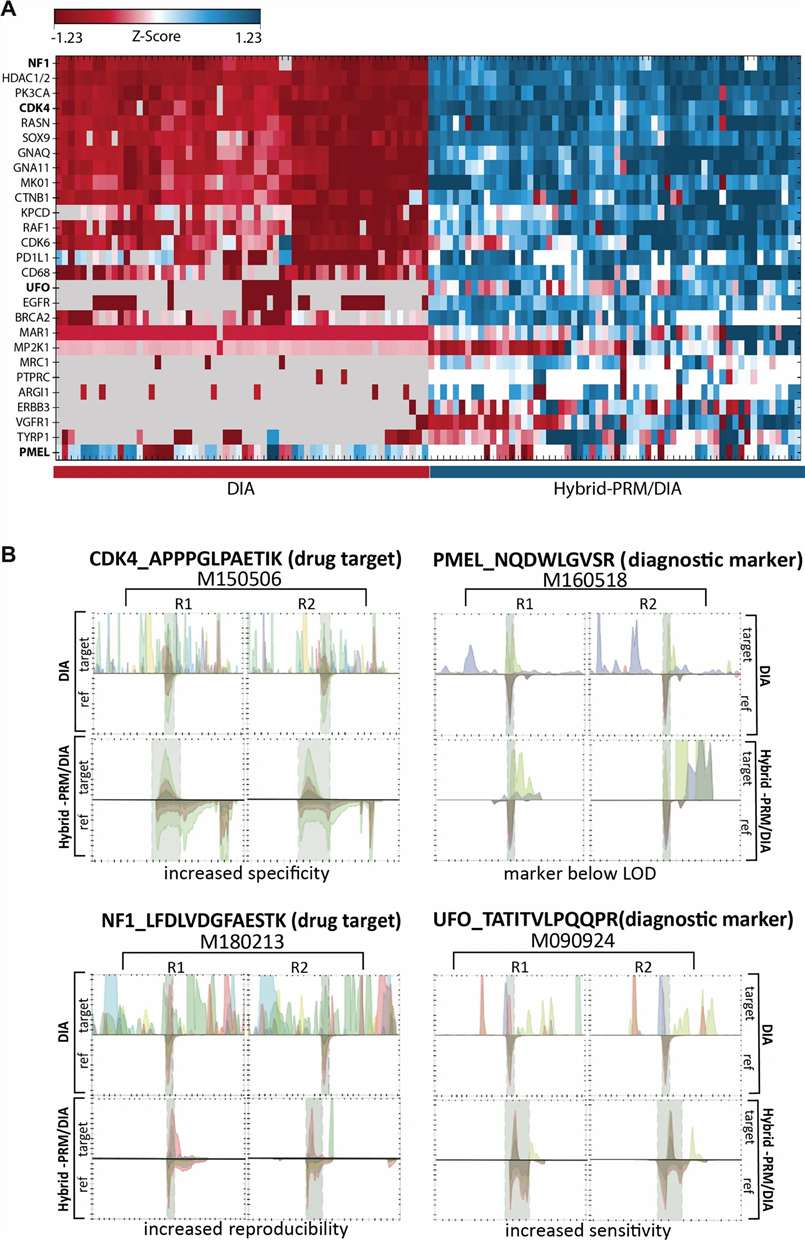
Hybrid PRM/DIA combines targeted biomarker monitoring with broad DIA-scale proteome coverage in a single analytical workflow.
Project Significance
This study shows that broad discovery-scale profiling and focused target monitoring do not need to be separated into disconnected analytical phases. By integrating PRM and DIA within one workflow, the method supports both comprehensive proteome digitization and more sensitive monitoring of predefined candidate biomarkers from the same sample context.
For translational proteomics projects, this type of workflow is particularly relevant when moving from early candidate screening toward downstream verification while preserving workflow continuity and minimizing additional sample consumption.
Reference
Goetze, S. et al. "Simultaneous targeted and discovery-driven clinical proteotyping using hybrid-PRM/DIA." Clinical Proteomics 21, 26 (2024). https://doi.org/10.1186/s12014-024-09478-5

