CORE SERVICE
Unlock Actionable Biomarkers
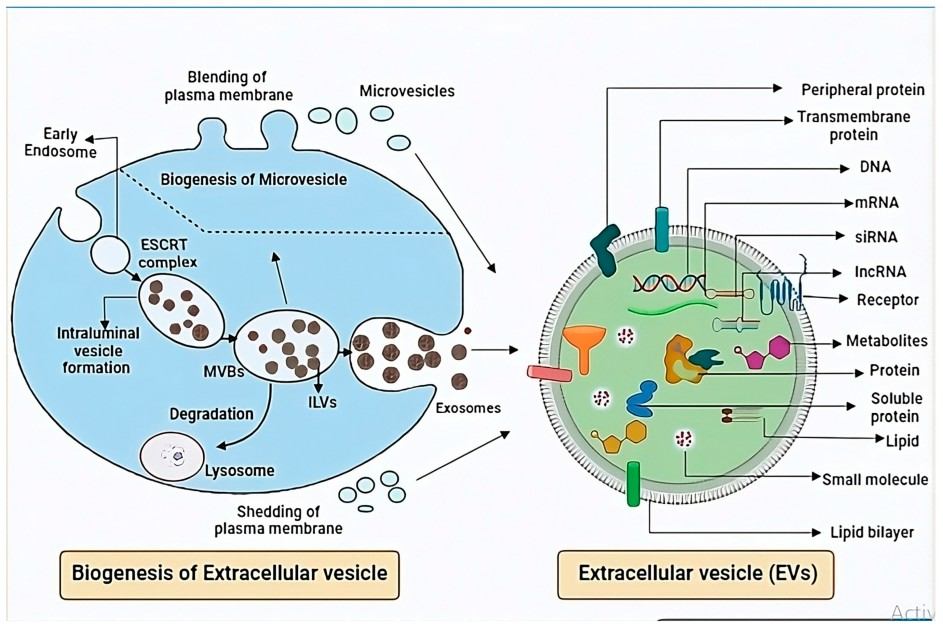
Unlock actionable biomarkers hidden in exosomes. Our Exosome Proteomics Services deliver high-depth, quantitative protein profiling from complex biofluids, enabling reliable biomarker discovery and mechanistic insights even from low-input samples. Backed by advanced mass spectrometry and proven workflows, we help researchers generate reproducible data that supports confident biological conclusions.
- High-depth detection: DIA and 4D-DIA workflows capture low-abundance exosomal proteins with robust quantification
- Flexible strategies: Label-free, labeled, PTM, and targeted PRM/MRM analyses tailored to your study goals
- Authoritative analysis: Advanced bioinformatics and expert interpretation grounded in published standards and large-scale project experience