Comprehensive glycosylation characterisation of glycoproteins is required by the ICH Q6B guidelines. For thorough structural characterisation, Creative Proteomics produces all the data required. If there are numerous glycosylation sites present, this is true for both the overall glycosylation population and the glycosylated structure of a single glycosylation site.
Why do N-linked Glycosylation Analysis?
N-linked glycosylation is a type of protein modification that involves the attachment of sugar molecules, specifically an oligosaccharide, to asparagine residues in proteins. This modification occurs in the endoplasmic reticulum and is essential for the proper folding, stability, and function of many proteins.
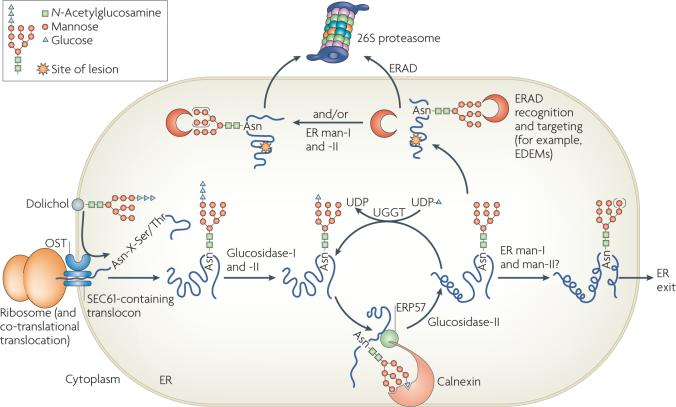 Fig 1. N-linked glycosylation and the degradation of glycosylated proteins. (Vembar, S. S., & Brodsky, J. L.; 2008)
Fig 1. N-linked glycosylation and the degradation of glycosylated proteins. (Vembar, S. S., & Brodsky, J. L.; 2008)
The process of N-linked glycosylation involves the following steps:
1. Synthesis of the oligosaccharide precursor: The oligosaccharide precursor, consisting of 14 sugar residues, is assembled on a lipid carrier called dolichol phosphate in the endoplasmic reticulum membrane.
2. Transfer of the oligosaccharide to the protein: The assembled oligosaccharide is transferred en bloc from dolichol phosphate to the amide nitrogen of an asparagine residue within the consensus sequence Asn-X-Ser/Thr, where X can be any amino acid except proline.
3. Protein folding and quality control: N-linked glycosylation can occur co-translationally or post-translationally. During protein synthesis, the glycosylation process can assist in proper protein folding. Misfolded proteins are targeted for degradation.
4. Glycan trimming and processing: The attached oligosaccharide undergoes trimming and modification in the endoplasmic reticulum and Golgi apparatus. Enzymes sequentially remove specific sugar residues and add others to produce a diverse range of glycan structures.
5. Biological functions: N-linked glycosylation plays various roles in protein function and cellular processes. It can affect protein stability, solubility, trafficking, and interactions with other molecules. Additionally, glycosylation is involved in cell-cell recognition, immune responses, and pathogen-host interactions.
What can we offer for N-linked Glycosylation Analysis?
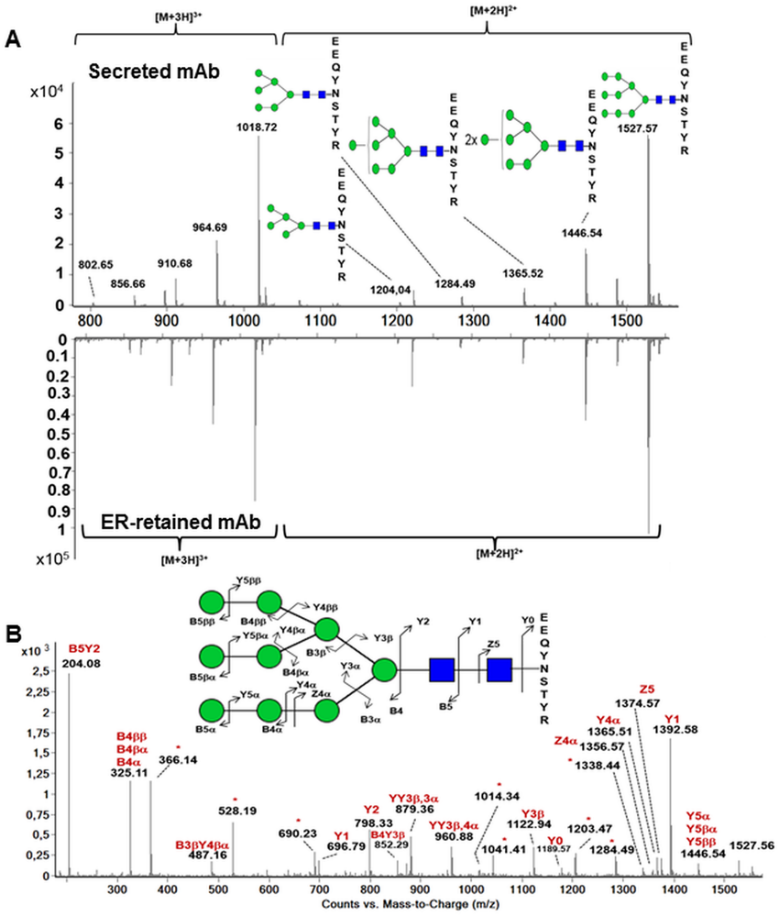 Fig 2. Analysis of the N-glycosylation of the heavy chain by mass spectrometry. (Vanier, G., et al.; 2015)
Fig 2. Analysis of the N-glycosylation of the heavy chain by mass spectrometry. (Vanier, G., et al.; 2015)
MS is a widely used technique for the analysis of glycosylation. It allows for the determination of the mass and structure of glycans attached to proteins. Different MS-based techniques, such as MALDI-TOF MS and ESI-MS, can be employed for glycosylation analysis.
- Liquid chromatography (LC)
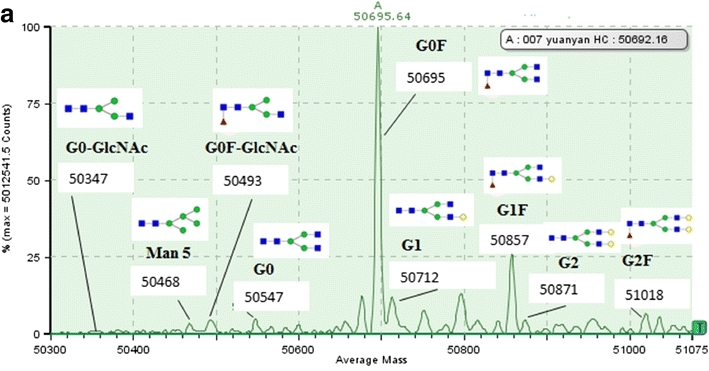 Fig 3. Analysis of N-linked glycosylation by LC-MS (Miao, S., et al.; 2017)
Fig 3. Analysis of N-linked glycosylation by LC-MS (Miao, S., et al.; 2017)
LC can be combined with MS or other detection methods, such as fluorescence or UV detection, to separate and analyze glycosylated proteins. Various LC techniques, including size exclusion chromatography, ion exchange chromatography, and affinity chromatography, can be used for glycosylation analysis.
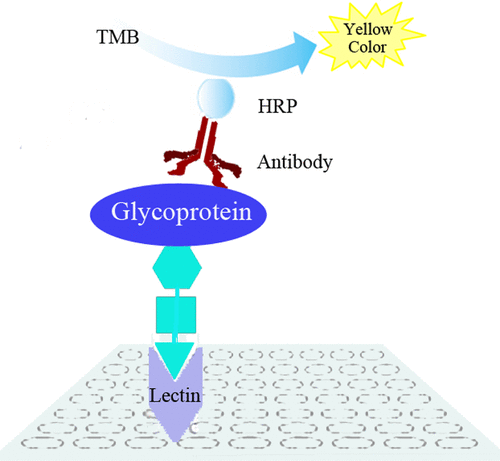 Fig 4. Diagram of reverse lectin-based ELISA assay for the analysis of glycosylation of target glycoproteins. (Wu, J., et al.; 2014)
Fig 4. Diagram of reverse lectin-based ELISA assay for the analysis of glycosylation of target glycoproteins. (Wu, J., et al.; 2014)
Lectins are proteins that specifically bind to carbohydrates. By using lectins with specific binding properties, such as Concanavalin A (ConA) or Wheat germ agglutinin (WGA), the presence and type of glycosylation can be detected.
- Enzymatic deglycosylation
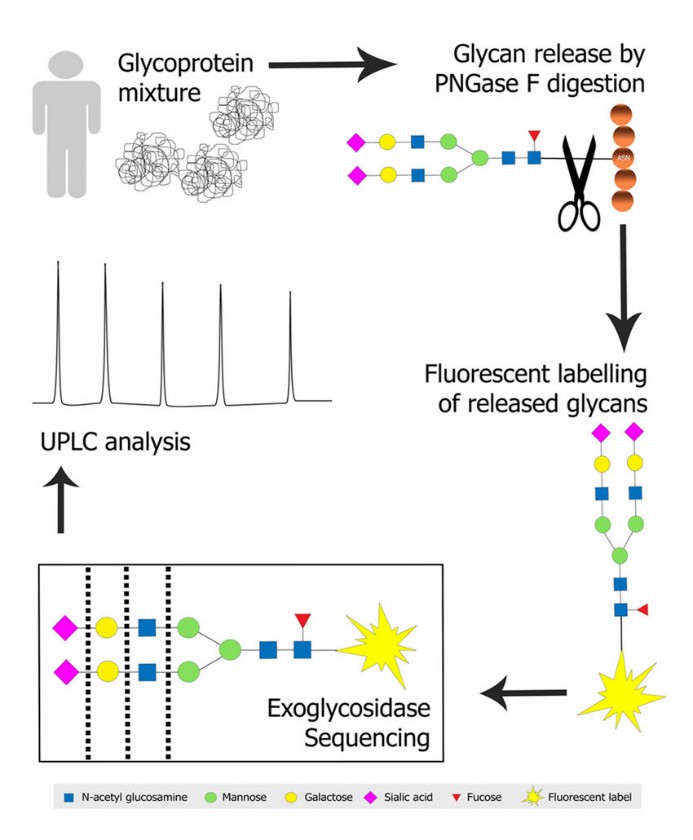 Fig 5. N-glycan profiling of a glycoprotein mixture. (Regan, P., et al.; 2019)
Fig 5. N-glycan profiling of a glycoprotein mixture. (Regan, P., et al.; 2019)
Specific enzymes, such as peptide N-glycosidase F (PNGase F), can be used to remove N-linked glycans from proteins. The resulting deglycosylated proteins can be analyzed by various methods, such as SDS-PAGE or western blotting.
- Nuclear magnetic resonance (NMR) spectroscopy
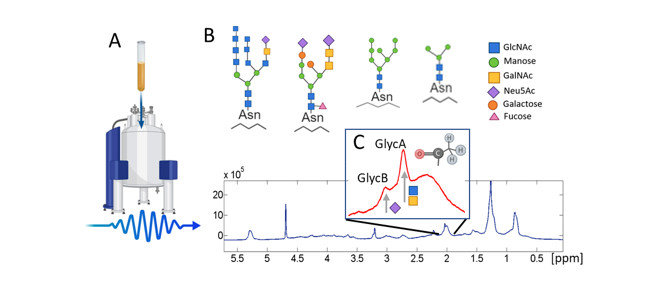 Fig 6. 1 H-NMR (nuclear magnetic resonance) glycoprotein analysis methodology. (Fuertes-Martín, et al.; 2019)
Fig 6. 1 H-NMR (nuclear magnetic resonance) glycoprotein analysis methodology. (Fuertes-Martín, et al.; 2019)
NMR spectroscopy can provide information about the conformational preferences of glycan structures attached to proteins.
Workflow for N-linked Glycosylation Analysis
The released N-linked glycan is purified, then separated by LC or CE after being labeled at the reducing end with an appropriate fluorescent labeling reagent. The N-linked glycan is then excited after peaks in the chromatogram or electropherogram are allocated using established databases and/or future experiments.
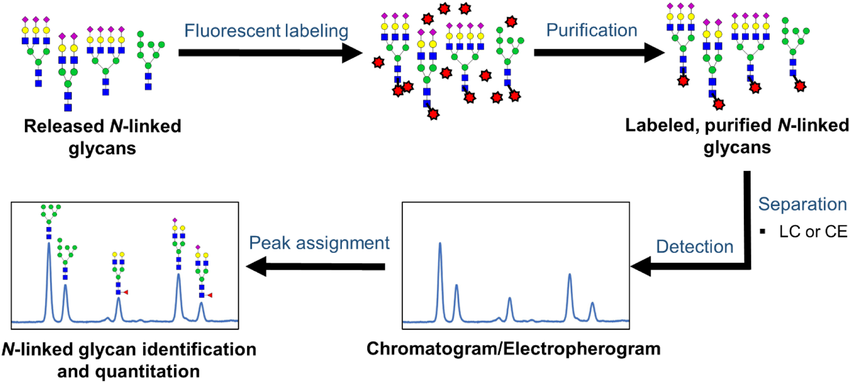 Fig 7. Workflow for the analysis of released N‐linked glycans via LC/CE‐fluorescence detection. (Patabandige, M. W., et al.; 2021)
Fig 7. Workflow for the analysis of released N‐linked glycans via LC/CE‐fluorescence detection. (Patabandige, M. W., et al.; 2021)
Our Benefits
Want to Learn More?
As a leader in proteomics and bioinformatics solutions, Creative Proteomics offers a wide range of services for protein characterization. With our advanced technologies and rigorous quality control measures, we ensure accurate and reliable identification and characterization of proteins. If you are interested in our service, please contact us for more details.
References
- Vembar, S. S., & Brodsky, J. L. One step at a time: endoplasmic reticulum-associated degradation. Nature Reviews Molecular Cell Biology. 2008, 9(12), 944–957.
- Vanier, G., et al.; Biochemical Characterization of Human Anti-Hepatitis B Monoclonal Antibody Produced in the Microalgae Phaeodactylum tricornutum. PLOS ONE. 2015, 10(10), e0139282.
- Miao, S., et al.; Identification of multiple sources of the acidic charge variants in an IgG1 monoclonal antibody. Applied Microbiology and Biotechnology. 2017, 101(14), 5627–5638.
- Wu, J., et al.; Analysis of Glycan Variation on Glycoproteins from Serum by the Reverse Lectin-Based ELISA Assay. Journal of Proteome Research. 2014, 13(4), 2197–2204.
- Regan, P., et al.; Early Stage Glycosylation Biomarkers in Alzheimer’s Disease. Medicines. 2019, 6(3), 92.
- Fuertes-Martín, et al.; Title: Human Serum/Plasma Glycoprotein Analysis by 1H-NMR, an Emerging Method of Inflammatory Assessment.Journal of Clinical Medicine. 2020, 9(2), 354.
- Patabandige, M. W., et al.; Quantitative clinical glycomics strategies: A guide for selecting the best analysis approach. Mass Spectrometry Reviews. 2021.
Related Sections
Services
Applications
For research use only, not intended for any clinical use.

 Fig 1. N-linked glycosylation and the degradation of glycosylated proteins. (Vembar, S. S., & Brodsky, J. L.; 2008)
Fig 1. N-linked glycosylation and the degradation of glycosylated proteins. (Vembar, S. S., & Brodsky, J. L.; 2008) Fig 2. Analysis of the N-glycosylation of the heavy chain by mass spectrometry. (Vanier, G., et al.; 2015)
Fig 2. Analysis of the N-glycosylation of the heavy chain by mass spectrometry. (Vanier, G., et al.; 2015) Fig 3. Analysis of N-linked glycosylation by LC-MS (Miao, S., et al.; 2017)
Fig 3. Analysis of N-linked glycosylation by LC-MS (Miao, S., et al.; 2017) Fig 4. Diagram of reverse lectin-based ELISA assay for the analysis of glycosylation of target glycoproteins. (Wu, J., et al.; 2014)
Fig 4. Diagram of reverse lectin-based ELISA assay for the analysis of glycosylation of target glycoproteins. (Wu, J., et al.; 2014) Fig 5. N-glycan profiling of a glycoprotein mixture. (Regan, P., et al.; 2019)
Fig 5. N-glycan profiling of a glycoprotein mixture. (Regan, P., et al.; 2019) Fig 6. 1 H-NMR (nuclear magnetic resonance) glycoprotein analysis methodology. (Fuertes-Martín, et al.; 2019)
Fig 6. 1 H-NMR (nuclear magnetic resonance) glycoprotein analysis methodology. (Fuertes-Martín, et al.; 2019) Fig 7. Workflow for the analysis of released N‐linked glycans via LC/CE‐fluorescence detection. (Patabandige, M. W., et al.; 2021)
Fig 7. Workflow for the analysis of released N‐linked glycans via LC/CE‐fluorescence detection. (Patabandige, M. W., et al.; 2021)
