- Service Details
- Case Study
What is Glycan Linkage Analysis?
Glycans, often referred to as carbohydrates or saccharides, consist of sugar molecules intricately connected. The primary building blocks are monosaccharides, and the linkages between them create glycan structures. These structures can be linear or branched, forming a complex array of carbohydrate configurations.
Glycan linkage analysis is a sophisticated analytical technique designed to unravel the intricate connections between monosaccharides within glycan structures. As glycans are complex carbohydrates composed of sugar molecules, understanding how these sugars are linked together provides critical insights into their functions and biological significance.
At its core, glycan linkage analysis involves determining the specific bonds and spatial arrangements between individual sugar units in a glycan molecule. This process often requires the use of advanced analytical methods such as mass spectrometry, nuclear magnetic resonance spectroscopy, and chromatographic techniques. By dissecting the glycan structure at the linkage level, researchers gain a nuanced understanding of the glycome, the complete set of glycans within a biological system.
Glycan linkage analysis is crucial in biological and medical research, revealing the roles of glycans in cellular processes like signaling, immune response, and protein folding. This analysis deciphers specific sugar arrangements, unlocking the language of cellular communication. Glycans, acting as molecular signals, influence key cellular events such as adhesion and recognition. In diseases like cancer, aberrant glycan linkages are common, and analyzing them helps identify unique glycan signatures. This insight enhances diagnostics and facilitates personalized treatment strategies, advancing medical research and patient care.
What We Can Provide
Glycan Linkage Position Analysis: Examines the specific positions of linkages within glycan structures, enhancing understanding at the molecular level.
Glycan Linkage Type Analysis: Focuses on identifying the types of linkages present, crucial for decoding the functional significance of glycans.
Glycan Branching Analysis: Explores the branching patterns within glycan structures, offering detailed information on their complexity and arrangement.
Glycan Cross-linking Analysis: Investigates cross-linkages between different glycan molecules, providing a holistic view of intermolecular connections.
Technology Platform for Glycan Linkage Analysis
Linkage analysis at Creative Proteomics is instrumental in determining the identities of diverse linkages between monosaccharides on glycoproteins and carbohydrates. This technology platform aims to provide a detailed characterization of glycans by unveiling specific linkages, enhancing our understanding of functional importance. The synergy with oligosaccharide population analysis contributes to a comprehensive view of glycan structures. Creative Proteomics employs advanced techniques, including mass spectrometry and chromatographic methods, ensuring a unified and thorough analysis for unraveling the complexities of glycan structures.
- Mass Spectrometry (MS): MALDI-TOF MS and ESI-MS/MS techniques deliver detailed insights into glycan structures. MS is versatile, suitable for various sample types, ensuring accurate molecular weight determination.
- Nuclear Magnetic Resonance (NMR) Spectroscopy: NMR spectroscopy plays a vital role in atomic-level glycan structure analysis. Leveraging NOESY and HSQC techniques enhances resolution and accuracy. NMR is applicable to diverse sample types, offering insights into spatial arrangements and specific linkages.
- Chromatographic Methods: High-Performance Liquid Chromatography (HPLC) is a versatile tool, utilizing techniques like HILIC and PGC for separation based on physicochemical properties. With applicability to various sample types, HPLC enables in-depth analysis of glycan linkages in complex mixtures.
- Capillary Electrophoresis (CE): Gaining prominence for high resolution and sensitivity, CE techniques like CGE and CE-MS facilitate effective separation and characterization. Adaptable to diverse sample matrices, CE is suitable for the analysis of complex samples.
Applications of Glycan Linkage Analysis
Biomedical Research: Unraveling the complexities of glycan structures aids biomedical research, providing crucial insights into diseases like cancer. The identification of unique glycan signatures serves as potential biomarkers for diagnostics and therapeutic target discovery.
Pharmaceutical Development: Ensuring the quality control of therapeutic glycoproteins, glycan linkage analysis is indispensable in pharmaceutical development. It guarantees the consistency and safety of biologic products, meeting rigorous regulatory standards.
Disease Biomarker Discovery: Glycan linkage analysis plays a key role in the discovery of disease-specific biomarkers. Analyzing glycan structures in biological samples helps identify patterns associated with various diseases, enhancing diagnostic precision and prognostic capabilities.
Immunology and Vaccine Development: In the realm of immunology, glycan linkage analysis contributes significantly to understanding immune responses. Deciphering glycan epitopes recognized by the immune system is crucial for designing vaccines targeting specific glycan structures, advancing vaccine development against infectious diseases.
Structural Biology: Providing essential information about the three-dimensional arrangement of glycans, glycan linkage analysis is fundamental in structural biology. This knowledge enhances our understanding of cell surface interactions, host-pathogen dynamics, and other intricate biological processes.
Sample Requirements for Glycan Linkage Analysis
| Sample Type | Recommended Sample Size |
|---|---|
| Serum or Plasma | 10-100 µL |
| Tissue Biopsy | 5-20 mg |
| Glycoprotein | 1-5 µg |
| Cell Culture Medium | 1 mL |
Case. Comprehensive Analysis of Heteroglycan Composition and Enzyme Activities in Pea Seedlings
Background:
This study aims to understand the heteroglycan composition and enzyme activities in pea seedlings (Pisum sativum L. 'Argenteum' mutant). The investigation focuses on protoplasts and chloroplasts, exploring their isolation processes and associated biochemical analyses.
Samples:
Pea seedlings were grown under controlled conditions, and various plant materials, including leaflets for protoplast isolation and intact chloroplasts, were harvested within the first 2 h of the light period. Soluble heteroglycan (SHG) preparations from pea leaflets were also analyzed.
Technical Method:
Protoplast Isolation:
- Modified procedure based on Edwards et al. (1978).
- Incubation in MES-buffered mannitol.
- Treatment with a mixture containing pectinase, cellulase, BSA, and PVP.
- Separation, washing, and fractionation into non-pelletable and pelletable fractions.
Chloroplast Isolation:
- Intact chloroplasts isolated within the first 2 h of the light period.
- Purification through centrifugation with Percoll.
- Confirmation of intactness via ferricyanide-dependent oxygen evolution.
Soluble Heteroglycan Preparation:
- Freezing of pea leaflets, extraction with ethanol.
- Isolation of protoplasts or chloroplasts.
- Centrifugation, heating, and lyophilization to obtain SHG preparations (SHGT, SHGS, SHGL).
Analytical Techniques:
- Non-denaturing PAGE and SDS-PAGE for protein separation.
- Assays for phosphorylase and glycan elongation activities.
- Precipitation of glycans with β-glucosyl Yariv reagent.
- Glycosidic linkage analysis through permethylation and derivatization.
Quantification and Analysis:
- Standard assays for protein, chlorophyll, and carbohydrate quantification.
- Ion chromatography (HPAEC-PAD) for monosaccharide and oligosaccharide separation.
- Field flow fractionation and multiangle laser light scattering for analyzing SHGL.
Results:
The study provides a comprehensive analysis of pea seedlings, revealing insights into the heteroglycan composition, enzyme activities, and the expression of Pho 2. Key findings include the characterization of SHG fractions, quantification of proteins and chlorophyll, and detailed insights into glycosidic linkages in the investigated samples.
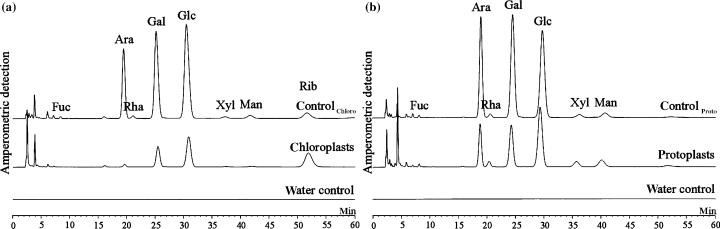 Monomeric composition of SHGT analysed by HPAEC-PAD.
Monomeric composition of SHGT analysed by HPAEC-PAD.
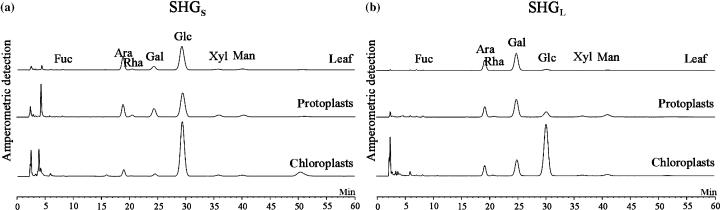 Monomeric composition of the SHGT-derived subfractions analysed by HPAEC-PAD.
Monomeric composition of the SHGT-derived subfractions analysed by HPAEC-PAD.
Reference
- Fettke, Joerg, et al. "The glycan substrate of the cytosolic (Pho 2) phosphorylase isozyme from Pisum sativum L.: identification, linkage analysis and subcellular localization." The Plant Journal 39.6 (2004): 933-946.




