- Service Details
- Case Study
What is Glycolipid?
Glycolipids are a type of lipid that contain a carbohydrate (sugar) moiety attached to a lipid molecule. They are important components of cell membranes and play key roles in cell-to-cell recognition, cell signaling, and membrane stability. Structurally, glycolipids consist of a hydrophobic lipid tail (such as a fatty acid or sphingosine) linked to a hydrophilic carbohydrate head group.
Glycolipids are diverse and may differ in structure, composition, and function depending on the specific types of carbohydrate and lipid components. Common classes of glycolipids include cerebrosides, gangliosides, and sulfatides. These molecules are present in various tissues and cell types throughout the body and are involved in many biological processes, including neuronal development, immune response regulation, and cell adhesion.
By characterizing the composition and abundance of glycolipids in biological samples, researchers can gain insights into cellular processes, disease mechanisms, and potential therapeutic targets. Glycolipid profiling identifies biomarkers for diseases such as cancer, neurodegenerative diseases, and metabolic syndrome
In addition, glycolipid analysis contributes to our understanding of lipid metabolism, lipid signaling pathways, and the role of lipids in health and disease. Aids drug discovery and development by assessing the effects of drugs on glycolipid profiles and lipid-related pathways.
Creative Proteomics provides one-stop glycolipid analysis services tailored to meet the diverse needs of researchers around the world. If you have any questions or other needs, please contact us.
Glycolipid Analysis Services by Creative Proteomics
Glycolipid Composition Analysis
Creative Proteomics employs mass spectrometry to characterize glycolipid components in biological samples. High-resolution mass spectrometry combined with chromatographic techniques identifies and quantifies various glycolipids, elucidating their molecular structure and relative abundance.
Glycolipid Analysis and Quantification
We provide glycolipid analysis and quantification services that enable you to gain a comprehensive understanding of the glycolipid profile of your biological samples. By employing both targeted and untargeted analytical strategies, we can accurately quantify the glycolipid species in different types of samples, thereby facilitating comparative studies and biomarker discovery efforts.
Glycolipid Structure Analysis
We use spectroscopic and chromatographic techniques to elucidate the molecular structure of glycolipids. Nuclear magnetic resonance (NMR) spectroscopy, mass spectrometry and liquid chromatography provide detailed structural information including fatty acid composition, linkage patterns and carbohydrate epitopes.
List of Detectable Glycolipid
| Gangliosides | Globosides | Sulfatides | Galactolipids | Glucosylceramide |
| Lactosylceramide | Ceramide trihexoside (CTH) | Cerebrosides | Sphingomyelin | Monogalactosyl diglyceride (MGDG) |
| Digalactosyl diglyceride (DGDG) | Sulfolipids | Glycoglycerolipids | Glycoproteolipids | Glycosphingolipids |
Technical Platforms for Glycolipid Analysis
Mass Spectrometry (MS)
Mass spectrometry has high sensitivity and specificity in identifying and quantifying molecules. Various MS techniques, including MALDI-TOF, ESI-MS, and tandem MS, enable comprehensive characterization of glycolipid structure and composition.
Liquid Chromatography (LC)
Liquid chromatography techniques such as HPLC and UPLC coupled with mass spectrometry can enhance the separation and analysis of glycolipids. LC-MS methods provide high-resolution analysis of complex glycolipid mixtures and facilitate detailed structural elucidation.
Nuclear Magnetic Resonance (NMR) Spectroscopy
Determine glycan linkage patterns, anomeric configurations, and lipid moieties. High-field NMR instruments enable precise characterization of glycolipid conformation and interactions.
Glycan Microarray
Glycan microarray technology enables high-throughput screening of glycolipid interactions with proteins, antibodies, and other biomolecules. This platform facilitates systematic analysis of glycolipid recognition and binding events and the discovery of novel glycolipid-protein interactions.
Bioinformatics and Data analysis
Bioinformatics tools facilitate data processing, structural annotation, and comparative analysis of glycolipid profiles. Advanced algorithms and databases simplify the interpretation of mass spectrometry and NMR data.
Sample Requirements for Glycolipid Analysis
| Sample Type | Sample Amount |
|---|---|
| Plasma | 100-200 μL |
| Serum | 100-200 μL |
| Tissue (e.g., liver, adipose) | 20-50 mg |
| Cell Culture | 1-5 x 10^6 cells |
| Urine | 1-2 mL |
| Cerebrospinal Fluid (CSF) | 50-100 μL |
Applications of Glycolipid Analysis
Biomarker Discovery: Glycolipids can serve as biomarkers for various diseases and conditions. Analyzing glycolipid profiles in biological samples such as blood, urine, or tissues can help identify potential biomarkers associated with diseases such as cancer, diabetes, and neurological disorders.
Understanding Disease Mechanisms: Studying glycolipids can provide insights into the underlying mechanisms of disease development and progression. Changes in glycolipid composition and metabolism have been linked to conditions like metabolic syndrome, neurodegenerative diseases, and infectious diseases.
Drug Discovery and Development: Glycolipid analysis can be used in drug discovery and development processes. By characterizing glycolipid profiles in response to drug treatments, researchers can identify potential therapeutic targets and evaluate the efficacy of candidate drugs.
Lipidomics Research: Glycolipid analysis is an integral part of lipidomics, the comprehensive study of lipid molecules in biological systems. Lipidomics approaches, including glycolipid profiling, are used to investigate lipid metabolism, lipid signaling pathways, and lipid-lipid interactions in health and disease.
Biotechnology and Biomedical Engineering: Glycolipids play essential roles in cell membrane structure, cell-cell recognition, and cell signaling. Analyzing glycolipids can contribute to the development of biomaterials, drug delivery systems, and tissue engineering approaches.
Clinical Diagnostics: Glycolipid analysis has clinical diagnostic applications, particularly in the diagnosis and monitoring of lipid-related disorders such as Gaucher disease, Fabry disease, and Tay-Sachs disease. Detecting abnormalities in glycolipid profiles can aid in early disease detection and management.
Nutritional Studies: Glycolipid analysis can be used to assess the impact of dietary interventions and nutritional factors on lipid metabolism and overall health. Understanding how glycolipid profiles change in response to diet can inform dietary recommendations and interventions for disease prevention and management.
Title: Bacterial Identification and Antimicrobial Resistance Detection through Glycolipid Mass Spectrometry
Background
Conventional diagnostic methods for bacterial infections often require days to weeks and may yield inaccurate results, necessitating the development of innovative technologies to improve patient outcomes. Glycolipid mass spectrometry offers a promising approach to bacterial identification and antimicrobial resistance detection by analyzing species-specific structural characteristics present in bacterial membranes.
Sample
The study utilized samples from various sources, including bacterial cultures grown in liquid media, polymicrobial mixtures containing multiple pathogens, and blood culture bottles inoculated with bacterial strains.
Technical Methods
Glycolipid Extraction: Bacteria were grown in liquid culture and subjected to a small-scale ammonium isobutyrate extraction to liberate glycolipids from polysaccharide constituents.
Mass Spectrometry Analysis: Extracts were analyzed using MALDI-TOF-MS in negative ion mode over a limited mass range (m/z 1,000–2,400) using norharmane as the matrix. Mass spectra were generated for individual strains, and reproducibility was evaluated through intra-/inter-day variability and technical extraction replicates.
Data Analysis: Mass spectra were processed, and signature ions unique to each species were identified. Pair-wise dot product analyses were conducted to measure similarity between mass spectra, enabling differentiation of bacterial species and detection of antimicrobial resistance markers.
Software Adaptation: The MALDI Biotyper software platform was adapted to recognize glycolipid mass spectra, allowing for species identification and distinguishing colistin-susceptible from resistant strains.
Clinical Application: The method was tested on polymicrobial mixtures and directly from blood culture bottles, demonstrating its applicability to complex clinical samples.
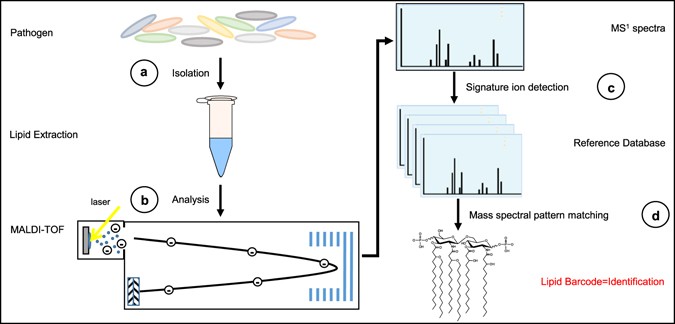 Strategy for glycolipid-based mass spectrometry platform for pathogen identification.
Strategy for glycolipid-based mass spectrometry platform for pathogen identification.
Results
The glycolipid mass spectrometry approach successfully differentiated bacterial species, detected antimicrobial resistance patterns, and identified pathogens directly from blood culture. It showed comparable accuracy to existing protein-based phenotyping methods and addressed their limitations, such as resistance detection and analysis of complex samples. Ongoing optimization efforts aim to further enhance the method's sensitivity and reduce processing time for clinical use.
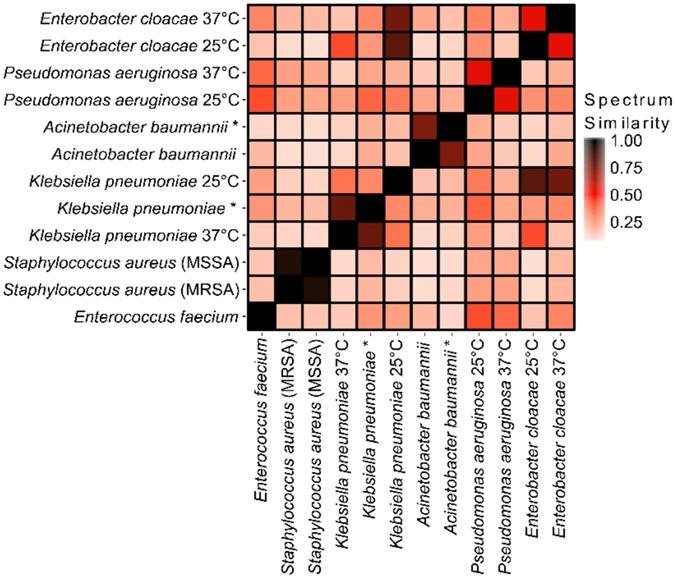 Dot product analysis of mass spectra for differentiation of ESKAPE pathogens.
Dot product analysis of mass spectra for differentiation of ESKAPE pathogens.
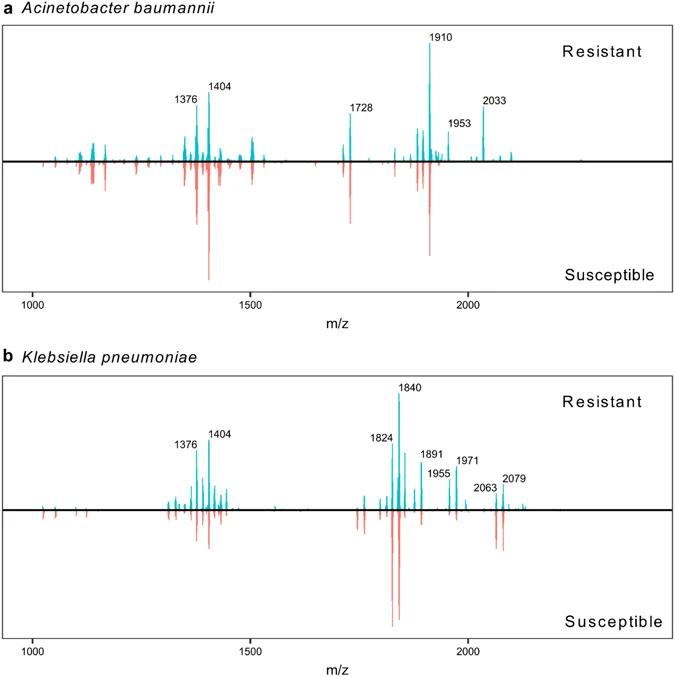 Consensus mass spectra were created by summation of detected ions from all replicates of colistin-resistant (top panels) and colistin-susceptible (bottom panels) A. baumannii (a) and K. pneumoniae (b).
Consensus mass spectra were created by summation of detected ions from all replicates of colistin-resistant (top panels) and colistin-susceptible (bottom panels) A. baumannii (a) and K. pneumoniae (b).
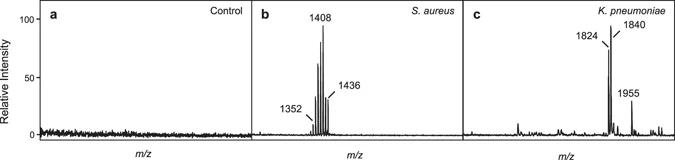 Detection of S. aureus and K. pneumoniae from blood culture.
Detection of S. aureus and K. pneumoniae from blood culture.
Reference
- Leung, Lisa M., et al. "Identification of the ESKAPE pathogens by mass spectrometric analysis of microbial membrane glycolipids." Scientific reports 7.1 (2017): 6403.




