- Service Details
- Case Study
What is Glycan Modification?
In the context of glycan structure analysis, modification analysis involves the examination and identification of various modifying groups or chemical alterations present in a polysaccharide molecule. Polysaccharides are complex carbohydrates composed of diverse sugar units, and their structural intricacies extend beyond the sugar backbone to include additional modifications.
The key aim of modification analysis is to elucidate the types, locations, and quantities of these modifications within the polysaccharide. This is pivotal for a comprehensive understanding of how these modifications impact the overall structure, function, and biological activities of the polysaccharide. Common modifications encompass processes such as methylation, phosphorylation, sulfation, and aldehydization, each contributing to the polysaccharide's unique characteristics.
To conduct modification analysis in glycan structures, researchers leverage sophisticated experimental techniques. Mass spectrometry allows for the determination of molecular weight, while nuclear magnetic resonance (NMR) spectroscopy provides insights into spatial arrangements. Chromatography techniques assist in separating and analyzing components, aiding in the identification of modifying groups.
What We Can Provide
Creative Proteomics offers comprehensive glycan modification analysis services, unraveling the complexities of glycan structures for advanced research. Our analysis covers a range of key modifications, including but not limited to:
Glycan Sulfation Modification Analysis: Explore sulfate group additions to glycan structures. Uncover how sulfation influences glycan-protein interactions and cellular processes.
Glycan Acetylation Modification Analysis: Delve into glycan acetylation. Gain insights into regulatory mechanisms affecting glycan stability and recognition.
Glycan Degradation Analysis: Discover glycan breakdown products. Understand glycan turnover and recycling's impact on cellular processes.
Glycan Methylation Modification Analysis: Unlock glycan methylation's regulatory roles. Explore how methylation influences glycan stability, signaling, and biological functions.
Technology Platform for Glycan Modification Analysis
Enzymatic Cleavage and Digestion:
Precise enzymatic cleavage is employed to selectively degrade glycan structures into smaller fragments. Enzymes such as glycosidases target specific glycosidic linkages, allowing for controlled degradation. This step is pivotal for obtaining a detailed profile of glycan modifications, as it reveals the positions and types of modifications within the polysaccharide.
Chromatography Techniques:
High-performance liquid chromatography (HPLC) and capillary electrophoresis (CE) are utilized for the separation and purification of glycans. HPLC, with various modes such as size-exclusion or ion-exchange, enables the isolation of glycan components based on size, charge, or other physicochemical properties. CE, on the other hand, excels in high-resolution separation of glycan isomers.
Mass Spectrometry (MS):
Mass spectrometry is a cornerstone technique providing high-resolution analysis of glycan structures. MALDI-TOF and ESI-MS allow for the precise determination of molecular weights, aiding in the identification of glycan modifications. Fragmentation techniques, such as tandem MS, offer insights into the structural details of modified glycans.
Nuclear Magnetic Resonance (NMR) Spectroscopy:
NMR spectroscopy offers detailed structural information, particularly regarding the spatial arrangements of atoms within glycan molecules. It is indispensable for elucidating glycosidic linkages, anomeric configurations, and the overall conformation of polysaccharides. High-resolution NMR allows for a comprehensive analysis of glycan modifications.
Fluorescent Labeling:
Fluorescent tags are strategically attached to specific glycan moieties. This labeling enhances sensitivity, enabling the visualization and quantification of modified glycans. Fluorescence detection is particularly useful in high-throughput analysis and glycan profiling studies.
Bioinformatics and Data Analysis:
Computational tools are employed for the interpretation of complex data generated by the analytical techniques. Bioinformatics algorithms aid in assigning glycan structures, identifying modification patterns, and correlating structural information with biological functions. This step ensures a comprehensive analysis and meaningful interpretation of the glycan modification data.
Applications of Glycan Modification Analysis
Biomedical Research:
- Disease Biomarkers: Glycan modifications are associated with various diseases, and analyzing these modifications can aid in identifying potential biomarkers for diseases such as cancer and neurodegenerative disorders.
- Glycoprotein Characterization: Studying glycan modifications on proteins helps understand their roles in cellular processes, immunity, and signaling pathways.
Pharmaceutical Development:
- Therapeutic Target Identification: Glycan analysis contributes to the identification of glycoproteins that may serve as therapeutic targets for drug development.
- Biopharmaceutical Quality Control: Monitoring glycan modifications is essential in ensuring the quality and consistency of glycoprotein-based biopharmaceuticals.
Glycoengineering:
- Bioproduction Optimization: Modifying glycan structures in cell lines used for bioproduction can enhance the efficiency of recombinant protein expression and improve therapeutic protein properties.
- Glycan Library Construction: Glycan modification analysis aids in the construction of glycan libraries, facilitating the study of glycan-protein interactions.
Immunology:
- Vaccine Development: Understanding glycan modifications on pathogens helps in designing vaccines that target specific glycan epitopes, enhancing immune responses.
- Autoimmune Diseases: Glycan analysis contributes to the investigation of glycan-related mechanisms in autoimmune diseases.
Glycomics and Systems Biology:
- Functional Glycomics: Glycan modification analysis is crucial for functional glycomics, providing insights into the roles of glycans in cell adhesion, migration, and communication.
- Systems Biology Integration: Integrating glycan data into systems biology approaches enhances our understanding of complex biological systems.
Food and Nutrition:
- Food Quality and Safety: Analyzing glycan modifications in food products can contribute to quality control and safety assessments.
- Nutritional Glycobiology: Investigating the impact of glycan modifications in dietary components on human health and nutrition.
Sample Requirements for Glycan Modification Analysis
| Sample Type | Sample Size Recommendation |
|---|---|
| Serum/Plasma | 100-500 μL |
| Cell Culture Supernatant | 1-5 mL |
| Tissue Homogenate | 10-50 mg |
| Glycoprotein/Glycopeptide | 1-10 μg |
| Biopharmaceuticals (e.g., mAbs) | 100-500 μg |
| Cerebrospinal Fluid (CSF) | 100-500 μL |
| Urine | 1-10 mL |
| Saliva | 100-500 μL |
| Food Products | Variable (depends on the matrix) |
Case. Characterization of Dual O-Acetylation in the Peptidoglycan of Lactobacillus plantarum and Its Functional Implications
Background:
The study focuses on the peptidoglycan (PG) structure of Lactobacillus plantarum, revealing the presence of two distinct types of O-acetylation—MurNAc O-acetylation and GlcNAc O-acetylation. This represents a unique feature, as it is the first bacterium known to harbor both forms of O-acetylation in its PG.
Samples:
The samples analyzed in the study are derived from L. plantarum NZ7100 (WCFS1 derivative), and the PG structure was determined through the analysis of muropeptides obtained after mutanolysin digestion.
Technical Method:
Analysis of PG Structure:
- RP-HPLC and MALDI-TOF mass spectrometry were employed to determine the PG structure of L. plantarum.
- Muropeptides obtained after mutanolysin digestion were analyzed for their m/z values, confirming the proposed structure.
Identification of O-Acetyltransferases:
- The BLASTP algorithm was used with the staphylococcal OatA protein as the query to identify O-acetyltransferases in L. plantarum.
- Two candidates, OatA (lp_0856) and OatB (lp_0925), were identified with distinct roles in MurNAc and GlcNAc O-acetylation, respectively.
Phylogenetic Analysis:
- Phylogenetic analysis revealed that OatA and OatB belong to different protein clusters, with OatA being closer to O-acetyltransferases in lactobacilli and lactococci.
Mutant Construction and PG Analysis:
- Deletion mutants for oatA and oatB were constructed, and a double oatA oatB mutant was also created.
- PG analysis of these mutants showed the specific roles of OatA and OatB in MurNAc and GlcNAc O-acetylation.
Functional Studies:
- Plate assays and PG digestion were performed to assess the impact of O-acetylation on lysozyme resistance.
- Autolysis tests and zymogram analysis were conducted to study the influence of O-acetylation on PGH (peptidoglycan hydrolase) activity, revealing differential effects on lysozyme and Acm2 N-acetylglucosaminidase.
Results:
Dual O-Acetylation in L. plantarum PG:
- L. plantarum is the first bacterium identified with both MurNAc and GlcNAc O-acetylation in its PG.
- The study provides a comprehensive characterization of the structural and functional aspects of these dual O-acetylations.
Identification of O-Acetyltransferases:
- Two O-acetyltransferases, OatA and OatB, were identified, each responsible for MurNAc and GlcNAc O-acetylation, respectively.
Functional Implications:
- MurNAc O-acetylation confers resistance to lysozyme and β-lactams, activating LytH amidase.
- GlcNAc O-acetylation inhibits Acm2 N-acetylglucosaminidase, modulating autolysis.
Evolutionary Insights:
- Evolutionary analysis suggests a horizontal gene transfer event for GlcNAc O-acetylation (oatB).
- L. plantarum's unique model demonstrates the co-evolution of synthesis/maturation PG machineries for both O-acetylation types.
Significance for PG Modification Control:
- L. plantarum's dual O-acetylation provides a unique model for studying the functional role of PG O-acetylation in controlling degradation and remodeling.
- The study highlights the plasticity of PG synthesis/degradation machineries and the importance of O-acetylation in regulating autolysins.
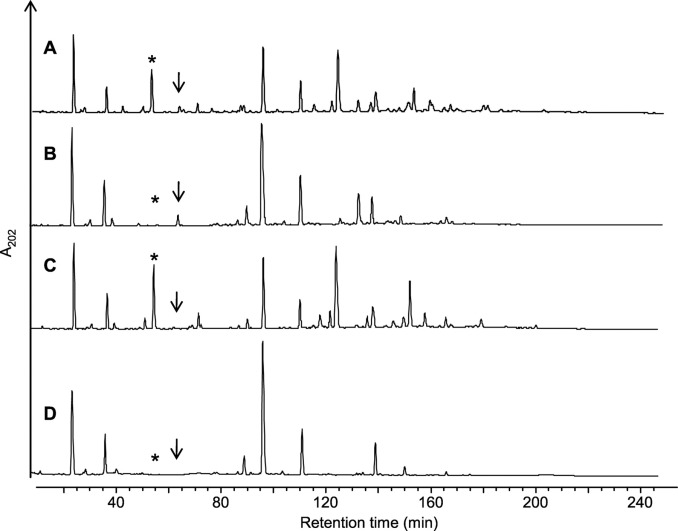 RP-HPLC separation of muropeptides from L. plantarum PG. WT (A), oatA mutant (B), oatB mutant (C), and oatA oatB mutant (D).
RP-HPLC separation of muropeptides from L. plantarum PG. WT (A), oatA mutant (B), oatB mutant (C), and oatA oatB mutant (D).
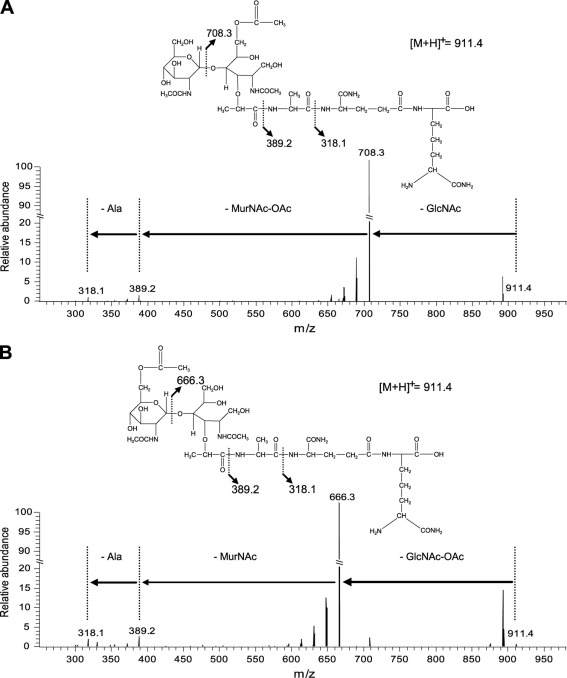 Fragmentation of O-acetylated dissacharide tripeptides by MS-MS.
Fragmentation of O-acetylated dissacharide tripeptides by MS-MS.
Reference
- Bernard, Elvis, et al. "Characterization of O-acetylation of N-acetylglucosamine: a novel structural variation of bacterial peptidoglycan." Journal of Biological Chemistry 286.27 (2011): 23950-23958.




