- Service Details
- Case Study
What is Glycoprotein Disulfide Bridge Mapping?
Glycoproteins, a diverse class of proteins adorned with carbohydrate chains, are ubiquitous in biological systems and play pivotal roles in various physiological processes. The structural integrity of glycoproteins is often maintained by disulfide bridges, covalent bonds formed between cysteine residues within the protein sequence. Understanding the precise arrangement of these disulfide bonds is crucial for deciphering the three-dimensional structure and functional properties of glycoproteins.
Glycoprotein disulfide bridge mapping involves the comprehensive analysis of glycoproteins to determine the exact connectivity of disulfide bonds within their structure. This technique integrates the high-resolution capabilities of liquid chromatography with the sensitive detection and mass analysis afforded by mass spectrometry, enabling the identification and characterization of disulfide linkages with remarkable precision.
Disulfide bridges contribute significantly to the stability, conformational rigidity, and functional diversity of glycoproteins. These intermolecular bonds are essential for maintaining proper protein folding, preventing misfolding and aggregation, and facilitating intra- and intermolecular interactions. Moreover, disulfide bonds can influence the biological activity, enzymatic function, and immunogenicity of glycoproteins, making them attractive targets for structural analysis.
Mapping the disulfide bridges of glycoproteins poses several challenges due to the complexity of their structures and the heterogeneity of their glycan attachments. Traditional methods for disulfide bond determination, such as X-ray crystallography and nuclear magnetic resonance (NMR) spectroscopy, may be limited by the size, solubility, and flexibility of glycoproteins. Additionally, the presence of carbohydrates can obscure the regions of interest or interfere with analytical techniques, necessitating specialized approaches for disulfide bridge mapping.
Creative Proteomics provides glycoprotein disulfide bridge mapping service by LC-MS technology.
Technology Platform for Glycoprotein Disulfide Bridge Mapping
High-Resolution Mass Spectrometry: Creative Proteomics employs state-of-the-art mass spectrometers with high-resolution capabilities, enabling accurate mass determination of peptides and their disulfide-linked species. This facilitates precise mapping of disulfide bridges within glycoproteins.
Chromatographic Separation: Glycoprotein digests are chromatographically separated based on their physicochemical properties, such as hydrophobicity, size, or charge. This step facilitates the isolation and purification of disulfide-linked peptides for subsequent mass spectrometric analysis.
Tandem Mass Spectrometry (MS/MS): Our platform utilizes tandem mass spectrometry techniques for fragmentation analysis of peptide ions, allowing for the sequencing of peptides and the identification of disulfide bond connectivity through the analysis of fragment ions.
Isotopic Labeling Strategies: Creative Proteomics employs isotopic labeling approaches such as iTRAQ (isobaric tags for relative and absolute quantitation) or SILAC (stable isotope labeling by amino acids in cell culture) to enable quantitative analysis of disulfide bridge patterns under different experimental conditions. This enhances our understanding of structural dynamics and protein interactions.
Bioinformatics Analysis: Our platform includes sophisticated bioinformatics tools and algorithms for data interpretation, including the identification of disulfide bond patterns, prediction of disulfide connectivity, and visualization of structural models. This enables comprehensive analysis and annotation of glycoprotein LC-MS data.
Specific Projects Offered by Creative Proteomics
- Structural Characterization: Accurate determination of disulfide bond connectivity within glycoproteins, providing crucial insights into their folding and stability.
- Comparative Analysis: Comparative profiling of disulfide bond patterns across different glycoprotein samples or experimental conditions, facilitating comparative proteomics studies.
- Quantitative Assessment: Quantitative analysis of disulfide-linked peptides to evaluate changes in disulfide bonding under various physiological or pathological conditions.
- Customized Solutions: Tailored experimental designs and analytical strategies to address specific research questions or challenges in glycoprotein characterization.
Sample Requirements for Glycoprotein LC-MS Disulfide Bridge Mapping
| Sample Type | Sample Amount |
|---|---|
| Biopsy (Tissue) | 5 - 20 mg |
| Bronchoalveolar Lavage Fluid (BALF) | 1 - 5 ml |
| Cell Culture Supernatant | 1 - 5 ml |
| Cell Lysate | 1 - 5 mg |
| CSF (Cerebrospinal Fluid) | 100 μl - 500 μl |
| Feces | 100 mg - 500 mg |
| Purified Glycoprotein | 50 μg - 100 μg |
| Saliva | 100 μl - 500 μl |
| Serum or Plasma | 100 μl - 500 μl |
| Synaptosomes | 50 - 100 μg |
| Synovial Fluid | 100 μl - 500 μl |
| Tissue Homogenate | 20 - 100 mg |
| Urine | 1 - 5 ml |
Case. Characterization of Sindbis Virus Structure Using Lysine Biotinylation and Mass Spectrometry
Background:
Sindbis virus, a prototype Alphavirus with a lifecycle involving vertebrates and mosquitoes, is utilized in various virus research fields. Despite previous structural studies, uncertainties exist regarding the accuracy of models derived from X-ray crystallography. Understanding the virus's structural features, particularly of the E2 protein, remains a challenge.
Samples:
Baby hamster kidney cells (BHK-21) infected with wild-type Sindbis virus were used for the study. Gradient-purified virus particles served as the primary sample for biotinylation and subsequent analysis.
Technical Method:
Gradient Purification and Particle/PFU Ratio Determination: Virus particles were purified using potassium tartrate step gradients, and particle/PFU ratios were calculated.
Biotinylation of Virus: Purified virus particles were labeled with sulfo-NHS-biotin at increasing molar excess concentrations, followed by quenching and titer determination.
Transmission Electron Microscopy and Negative Staining: Infected cell monolayers were prepared for transmission electron microscopy, and virus particles were directly attached to carbon-coated grids for negative staining.
Enzymatic Digestion: Biotinylated Sindbis virus was denatured, reduced, and digested with endoproteinase GluC or trypsin.
Monomeric Avidin Affinity Chromatography: Biotinylated peptides were affinity-purified using a monomeric avidin column.
Quadrupole Ion Trap LC-MS/MS: LC-MS/MS analysis was performed using an Agilent mass spectrometer coupled to an HPLC system, with spectral data analyzed using SpectrumMill software.
Results:
Effects of Biotinylation on Sindbis Virus Infectivity: Biotinylation at varying concentrations minimally affected virus infectivity until reaching a threshold, beyond which infectivity sharply decreased.
Identification of Biotinylation Sites: Lysine residues in the E1 and E2 glycoproteins were identified as biotinylated, providing insights into the virus's structural features.
Validation of Structural Models: Discrepancies between biotinylation data and X-ray crystallography models highlight limitations in existing structural models, particularly concerning the E1 glycoprotein's N-terminal region.
Enhanced Structural Insights: Biotinylation coupled with mass spectrometry facilitated detailed analysis of Sindbis virus structure, revealing surface-exposed lysine residues and antigenic regions.
Confirmation of Biotinylation Technique: The labeling process did not compromise virus infectivity, affirming the suitability of lysine biotinylation for structural analysis without perturbing viral functionality.
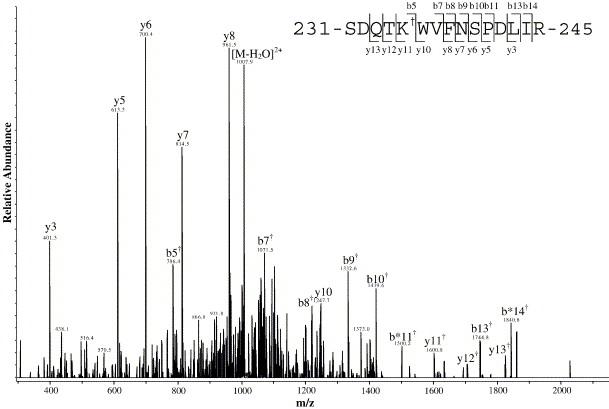 Representative MS/MS spectrum of a biotinylated peptide from the E2 glycoprotein.
Representative MS/MS spectrum of a biotinylated peptide from the E2 glycoprotein.
 Sites of biotinylation mapped onto the sequences of the E1 and E2 glycoproteins.
Sites of biotinylation mapped onto the sequences of the E1 and E2 glycoproteins.
Reference
- Sharp, Joshua S., et al. "Structural characterization of the E2 glycoprotein from Sindbis by lysine biotinylation and LC-MS/MS." Virology 348.1 (2006): 216-223.




