- Service Details
- Case Study
What is Protein Glycosylation?
Protein glycosylation is a vital post-translational modification process wherein carbohydrate molecules, known as glycans, are covalently attached to specific amino acid residues within a protein chain. This process occurs in all domains of life, from bacteria to humans, and plays crucial roles in various biological processes, including protein folding, stability, localization, and function.
There are two primary types of protein glycosylation:
- N-Linked Glycosylation: In N-linked glycosylation, glycans are attached to the nitrogen atom of asparagine (N) residues within the consensus sequence Asn-X-Ser/Thr, where X can be any amino acid except proline. This type of glycosylation predominantly occurs in the endoplasmic reticulum (ER) and Golgi apparatus and is involved in protein folding, quality control, and cellular recognition processes.
- O-Linked Glycosylation: O-linked glycosylation involves the attachment of glycans to the oxygen atom of serine (S) or threonine (T) residues in a protein chain. Unlike N-linked glycosylation, O-linked glycosylation lacks a defined consensus sequence and occurs predominantly in the Golgi apparatus and cytoplasm. O-glycans contribute to protein stability, protection against proteolysis, and cell-cell interactions.
Understanding the complex landscape of protein glycosylation is essential for elucidating the structure-function relationships of glycoproteins and their roles in health and disease. Protein glycosylation analysis enables we to:
- Identify glycosylation sites and characterize glycan structures.
- Investigate the heterogeneity and dynamics of glycan modifications.
- Determine the impact of glycosylation on protein structure, function, and stability.
- Discover novel glycan-protein interactions and therapeutic targets.
- Develop glycoprotein-based biomarkers for disease diagnosis and prognosis.
Services of Protein Glycosylation Analysis by Creative Proteomics
- N-Glycan Profiling: Identification and quantification of N-linked glycans attached to proteins.
- O-Glycan Analysis: Characterization of O-linked glycans attached to proteins.
- Glycoprotein Identification: Detection and identification of glycoproteins in complex mixtures.
- Glycosylation Site Mapping: Determination of specific amino acid residues where glycans are attached on proteins.
- Glycosylation Pattern Analysis: Studying the diversity and distribution of glycan structures on proteins.
- Glycosylation Quantitation: Measurement of glycan abundance and stoichiometry on proteins.
- Glycosylation Changes in Disease: Investigating alterations in protein glycosylation associated with various diseases.
- Glycosyltransferase Activity Assays: Evaluation of enzyme activity involved in glycan biosynthesis.
- Mass Spectrometry-based Glycoproteomics: Utilizing mass spectrometry techniques for comprehensive glycoprotein analysis.
Technology Platform for Protein Glycosylation Analysis
1. Sample Preparation: Extract proteins from the biological sample of interest (e.g., cell lysate, serum) and digest them into peptides using proteolytic enzymes (e.g., trypsin).
2. Glycopeptide Enrichment: Enrich glycopeptides from the peptide mixture using techniques such as lectin affinity chromatography or hydrazide chemistry. This step selectively isolates glycopeptides based on their glycan moieties.
3. LC-MS Analysis: Separate and analyze the enriched glycopeptides using liquid chromatography coupled with mass spectrometry (LC-MS). LC separates glycopeptides based on their hydrophobicity, while MS detects and characterizes the glycopeptides based on their mass-to-charge ratio (m/z).
4. Glycan Identification: Identify the glycan structures attached to the glycopeptides by analyzing their fragmentation patterns in MS/MS spectra. Software tools and databases are used to match the experimental data with known glycan structures.
5. Glycosylation Site Mapping: Determine the specific amino acid residues where glycans are attached on proteins by analyzing the MS/MS spectra of glycopeptides. This step helps map the glycosylation sites within the protein sequence.
6. Quantitative Analysis: Quantify the abundance of glycopeptides and their glycan structures using peak intensity or spectral counting methods. This allows for the comparison of glycosylation profiles between different samples or conditions.
Sample Requirements for Protein Glycosylation Analysis
| Sample Type | Sample Quantity | Notes |
|---|---|---|
| CSF (Cerebrospinal Fluid) | 100-500 µL | Collect using aseptic techniques. Avoid repeated freeze-thaw cycles. |
| Synovial Fluid | 100-500 µL | Collect using aseptic techniques. Avoid contamination with blood. |
| Feces | 100-500 mg | Collect in clean containers. Avoid contamination with urine or water. |
| Mucus | 100-500 µL | Collect using non-invasive techniques. Avoid contamination with saliva. |
| Tears | 100-200 µL | Collect using sterile techniques. Avoid contamination with eye drops. |
Applications of Protein Glycosylation Analysis
Biomedical Research: Understanding the roles of protein glycosylation in cellular processes, disease mechanisms, and therapeutic targets.
Biopharmaceutical Development: Characterization of glycosylation profiles of recombinant proteins for quality control and optimization of biologics production.
Diagnostic Biomarker Discovery: Identification of glycosylation-based biomarkers for disease diagnosis and prognosis.
Drug Development: Targeting glycan-mediated interactions for the development of novel therapeutics.
Food and Environmental Sciences: Analysis of glycoproteins in food products and environmental samples for safety and quality assessment.
Case. Advances in Quantification of Site-Specific Protein Glycoforms Using LC-MS/MS-MRM Techniques
Background:
Protein glycosylation, the addition of glycans to proteins, plays crucial roles in various biological processes and disease mechanisms. Understanding the site-specific glycosylation patterns of proteins is essential for elucidating their functions and identifying biomarkers for diseases. However, quantifying site-specific protein glycoforms with high sensitivity and specificity remains challenging due to the complexity of glycan structures and the dynamic nature of glycosylation.
Samples:
Complex biological samples, such as serum or protein digests, containing a mixture of proteins with diverse glycosylation patterns are commonly used for quantification studies. These samples often present challenges in terms of ion suppression and interference, necessitating enrichment techniques to isolate target glycopeptides for analysis.
Technical Method:
Fragmentation Analysis: The quantification of site-specific protein glycoforms relies on the fragmentation analysis of glycopeptides using LC-MS/MS-MRM techniques. Fragmentation patterns vary depending on the peptide sequence and attached glycan structure, with specific ions such as Y0 and m/z 138 commonly monitored for quantification.
Enrichment Strategies: Various enrichment strategies, including reverse glycobloting and 2D HPLC, are employed to isolate glycopeptides from complex samples. These techniques improve the sensitivity and specificity of quantification by reducing ion suppression and interference from co-eluting peptides.
Collision Energy Optimization: Structure-specific optimization of collision energy (CE) is crucial for maximizing sensitivity and specificity in quantification. Different glycan structures may require different CE settings to achieve optimal fragmentation patterns for accurate quantification.
MS3 Quantification: MS3 techniques involve monitoring abundant peptide fragments in addition to glycopeptide ions, offering increased sensitivity and specificity, especially for minor glycoforms. This approach enhances the reliability of quantification and enables the detection of low-abundance glycoforms in complex samples.
Challenges and Future Directions: Challenges in quantifying site-specific protein glycoforms include the lack of appropriate standards and the complexity of sample matrices. However, advancements in glycan and glycopeptide synthesis are expected to facilitate progress in glycoproteomics research, leading to more sensitive and specific quantification methods.
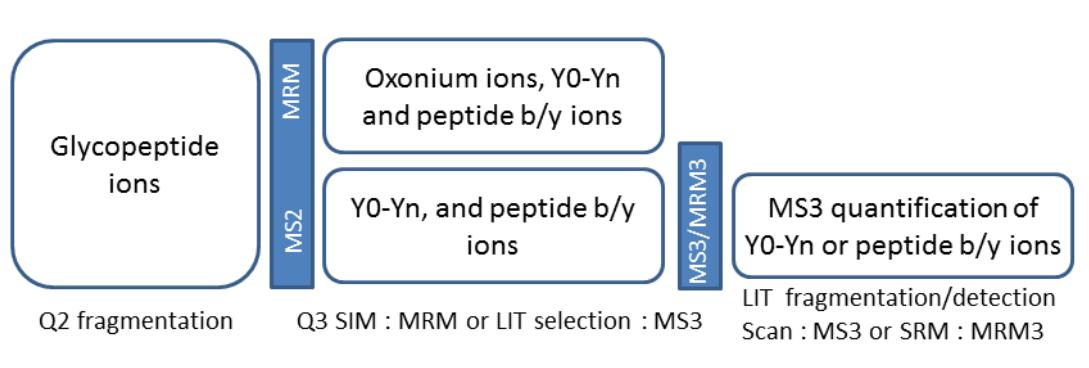 Schematic of MRM and MS3 workflows for quantification of glycopeptides.
Schematic of MRM and MS3 workflows for quantification of glycopeptides.
Results:
Overall, the reviewed studies demonstrate the feasibility of using LC-MS/MS-MRM techniques for sensitive and specific quantification of site-specific protein glycoforms. Enrichment techniques, fragmentation pattern analysis, optimization of CE, and MS3 quantification all contribute to improving the accuracy and reliability of glycoproteomics analyses. Despite challenges, continued advancements in methodology and technology are expected to further enhance the capabilities of LC-MS/MS-MRM for glycoform quantification in complex biological samples.
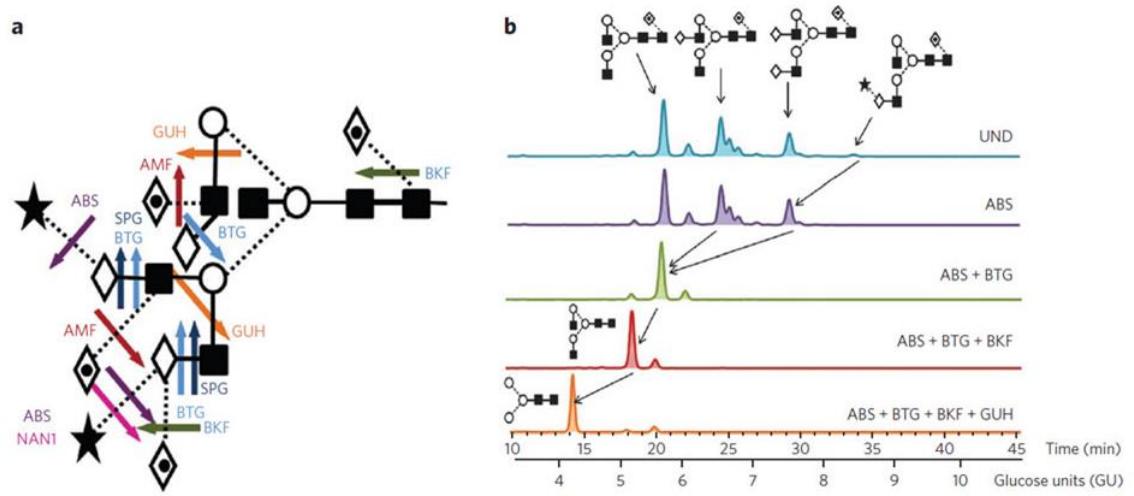 A. Cleavage sites of commonly used exoglycosidases for glycan fingerprinting. B. Representative HILIC profiles of 2-AB–labeled N-glycans from human IgG before (UND, undigested) and after exoglycosidase digestion.
A. Cleavage sites of commonly used exoglycosidases for glycan fingerprinting. B. Representative HILIC profiles of 2-AB–labeled N-glycans from human IgG before (UND, undigested) and after exoglycosidase digestion.
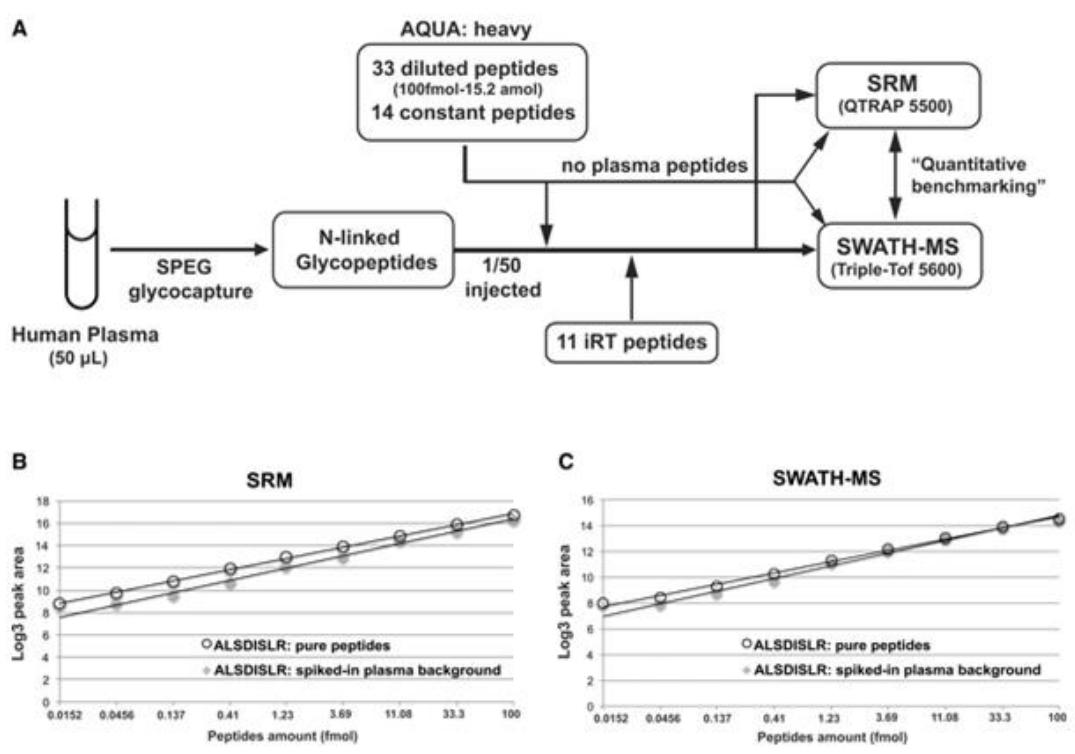 Quantitative measurements of N‐linked glycoproteins by SRM or SWATH‐MS.
Quantitative measurements of N‐linked glycoproteins by SRM or SWATH‐MS.
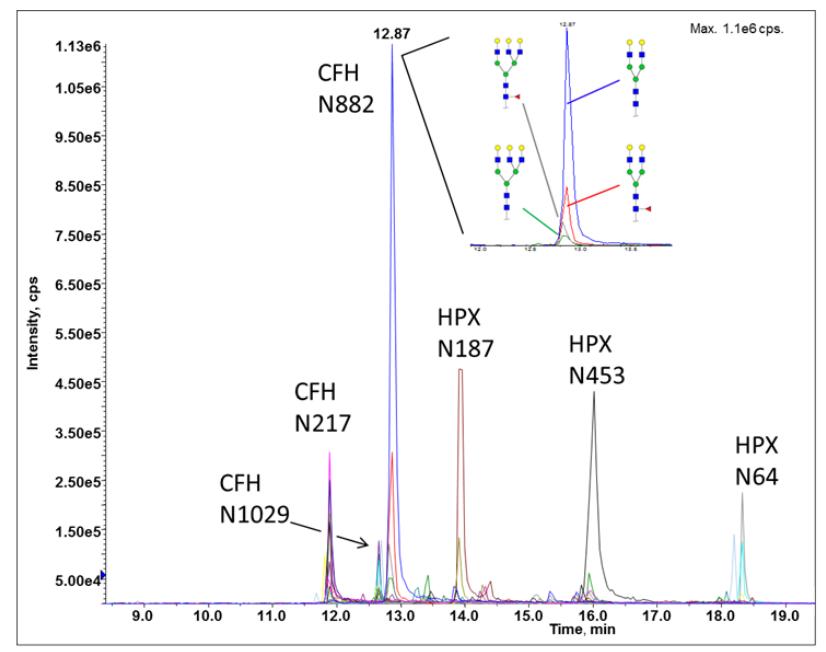 Site-specific glycoforms of sialidase treated hemopexin (HPX) and complement factor H (CFH) separated by RP chromatography and detected by MRM-MS
Site-specific glycoforms of sialidase treated hemopexin (HPX) and complement factor H (CFH) separated by RP chromatography and detected by MRM-MS
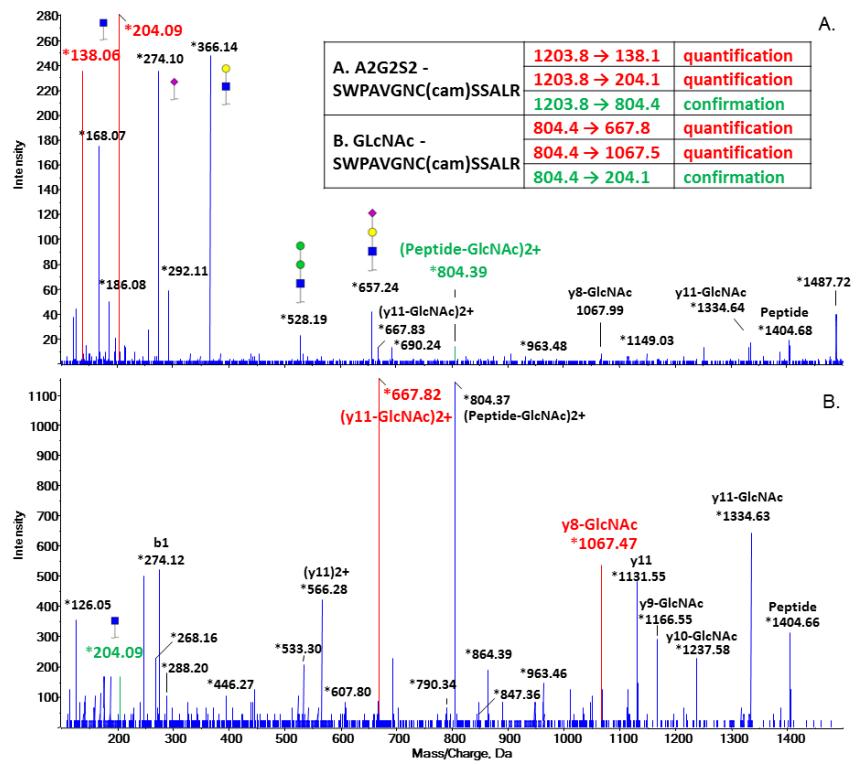 Selection of transitions for quantification and confirmation based on CID spectrum of A. biantennary sialylated glycopeptide (A2G2S2); B. GlcNAc-glycopeptide generated by Endo F3.
Selection of transitions for quantification and confirmation based on CID spectrum of A. biantennary sialylated glycopeptide (A2G2S2); B. GlcNAc-glycopeptide generated by Endo F3.
Reference
- Goldman, Radoslav, and Miloslav Sanda. "Targeted methods for quantitative analysis of protein glycosylation." PROTEOMICS–Clinical Applications 9.1-2 (2015): 17-32.




