- Service Details
- Case Study
What is N- and O-Linked Glycosylation Mapping?
N-linked and O-linked glycosylation mapping is a sophisticated analytical approach used in biological research to unveil the intricate patterns of carbohydrate modifications on proteins. N-linked glycosylation typically occurs at asparagine residues within the consensus sequence N-X-S/T (where X can be any amino acid except proline), while O-linked glycosylation predominantly targets serine or threonine residues. By mapping the sites of glycan attachment on proteins, we can gain valuable insights into protein structure, function, and regulation. This mapping process involves a combination of advanced techniques such as mass spectrometry, lectin affinity chromatography, glycosidase digestion, and chemical labeling, which collectively enable the precise identification and characterization of glycosylation sites. Through N- and O-linked glycosylation mapping, we can unravel the complex interplay between glycans and proteins, shedding light on their roles in health, disease, and biotechnological applications.
Creative Proteomics's N- and O-Linked Glycosylation Mapping Services
- N- and O-Glycan Profiling Service: To provide a complete picture of protein glycosylation, we offer a complete N- and O-glycan profiling service. This service not only identifies the glycans present, but also provides insight into their relative abundance in a given sample. Our service ensures the detection and analysis of both common and rare glycan species, thus helping to reveal hidden details in the glycosylation profile of proteins.
- N- and O-Glycosylation Site Analysis Services: We not only provide information on which amino acids are glycosylated, but also the precise sites of glycosylation. Understanding glycosylation at specific sites is critical because it directly affects protein folding, stability and activity. Our innovative use of technologies such as electron-transfer/higher-energy collision dissociation (EThcD) ensures the successful identification of these sites.
- N- and O-Glycan Linkage Analysis: Determination of the glycosidic linkages and branching patterns of N- and O-linked glycans, providing insights into glycan biosynthesis and structural diversity through enzymatic digestion, chromatographic separation, and mass spectrometry-based analysis.
- N- and O-Glycosylation Site Occupation: Quantitative assessment of glycosylation site occupancy on proteins, evaluating the extent to which specific amino acid residues are glycosylated under different physiological or experimental conditions, facilitating the study of glycan dynamics and protein glycosylation regulation.
Technology Platform for N- and O-Linked Glycosylation Mapping
Mass Spectrometry (MS):
Mass spectrometry lies at the core of our technology platform for glycosylation mapping. Our high-resolution mass spectrometers coupled with liquid chromatography enable the sensitive detection and characterization of glycopeptides, allowing for the precise localization of N- and O-linked glycosylation sites within protein sequences. By analyzing the mass spectra of glycopeptides, we can elucidate the composition and structure of glycans attached to proteins with exceptional accuracy.
Lectin Affinity Chromatography:
Leveraging the specificity of lectins for glycan structures, our platform includes lectin affinity chromatography methods for glycopeptide enrichment. Through selective binding to glycosylated proteins, lectins facilitate the isolation of glycopeptides from complex biological samples. Combined with mass spectrometry analysis, this approach enables the identification and quantification of N- and O-linked glycosylation sites, providing valuable insights into glycan diversity and protein glycosylation patterns.
Glycosidase Digestion:
Our technology platform encompasses glycosidase digestion strategies for the precise mapping of N- and O-linked glycosylation sites. By treating glycoproteins with specific glycosidases such as PNGase F for N-linked glycans and O-glycosidases for O-linked glycans, we can selectively remove glycans from protein sequences. Subsequent analysis by mass spectrometry allows for the identification of deglycosylated peptides, revealing the exact locations of glycosylation sites and elucidating the glycan occupancy of proteins.
Chemical Labeling Strategies:
In addition to enzymatic approaches, our platform offers innovative chemical labeling strategies for quantitative glycosylation mapping. Through the use of isotopically labeled reagents and advanced labeling techniques such as SILAC and iTRAQ, we enable comparative analysis of glycosylation patterns under different experimental conditions. This allows researchers to uncover changes in protein glycosylation profiles associated with disease states, drug treatments, or biological processes.
Sample Requirements for N- and O-Linked Glycosylation Mapping
| Sample Type | Sample Volume/Amount | Additional Notes |
|---|---|---|
| Cell Lysates | 1-5 mg | Suitable for global glycosylation profiling and site-specific glycosylation analysis. |
| Tissue Homogenates | 10-50 mg | Recommended for studying tissue-specific glycosylation patterns and disease biomarkers. |
| Serum/Plasma | 100-500 µL | Ideal for clinical glycoproteomics, biomarker discovery, and disease diagnosis. |
| Cerebrospinal Fluid (CSF) | 50-200 µL | Valuable for studying glycosylation changes associated with neurological disorders. |
| Urine | 1-5 mL | Useful for non-invasive biomarker discovery and monitoring of kidney-related diseases. |
| Saliva | 100-500 µL | Suitable for studying mucin-type glycoproteins and oral health-related biomarkers. |
| Recombinant Proteins | 10-100 µg | Important for glycoengineering studies, biopharmaceutical quality control, and protein optimization. |
| Glycoprotein Standards | 1-10 µg | Essential for method validation, calibration, and quality assurance in glycosylation analysis. |
| Cell Culture Supernatants | 1-5 mL | Useful for studying secreted glycoproteins, recombinant expression systems, and bioprocess monitoring. |
Case. Glycosylation Analysis of SARS-CoV-2 Variants Reveals Structural Adaptations
Background:
Understanding the glycosylation profile of the spike (S) protein of SARS-CoV-2 variants is crucial for deciphering their pathogenicity and immune evasion mechanisms. Glycosylation plays a pivotal role in viral-host interactions, and alterations in glycan structures can impact viral infectivity and immune recognition.
Samples:
The study analyzed recombinant S proteins of the SARS-CoV-2 variants Alpha, Beta, Gamma, Delta, Omicron, and the wild-type strain. These samples were expressed in HEK293F cells to mimic the glycosylation profile of the actual virus.
Technical Method:
1. Sample Preparation: Recombinant S proteins of SARS-CoV-2 variants (Alpha, Beta, Gamma, Delta, Omicron) and the wild-type strain were obtained from R&D Systems. These proteins were expressed in HEK293F cells to ensure similarity to the native viral glycosylation profile.
2. Protease Digestion: Multiple proteases, including trypsin, chymotrypsin, and α-lytic protease, were employed for site-specific glycosylation analysis. The recombinant S proteins were subjected to enzymatic digestion to cleave peptide bonds and release glycopeptides.
3. LC–MS/MS Analysis: The digested glycopeptides were analyzed using liquid chromatography-tandem mass spectrometry (LC–MS/MS). High-resolution mass spectrometry facilitated the identification and quantification of glycopeptides. HCD product-triggered CID tandem MS provided fragmentation patterns for glycopeptide characterization.
4. Data Analysis: LC–MS/MS data files were processed using Byonic software for glycopeptide annotation. A glycan database within Byonic was utilized for initial glycopeptide identification, followed by validation through manual inspection of spectra. Ambiguous spectra were excluded from analysis, ensuring accurate glycoform assignment.
5. Glycomic Profiling: Overall glycan distribution and composition were determined through glycomics analysis. Glycans were released from glycoproteins using appropriate enzymes (e.g., PNGase F for N-deglycosylation). Released glycans were separated from peptides using C18 solid-phase extraction (SPE) and analyzed by LC–MS/MS.
6. Quantification and Interpretation: Quantitative analysis of glycoforms at each glycosylation site was performed by evaluating precursor MS1 peak areas. Relative abundances of glycoforms were calculated manually using specialized software (e.g., Xcalibur, FreeStyle) after spectral deconvolution. The significance of glycosylation changes was interpreted in the context of viral evolution, transmissibility, and immune evasion mechanisms.
7. Statistical Analysis: Statistical methods, such as principal component analysis (PCA) and heatmap visualization, were employed to compare glycosylation profiles across variants and the wild-type strain. These analyses helped identify significant glycosylation differences and patterns associated with viral adaptation and pathogenicity.
8. Quality Control: Stringent quality control measures were implemented throughout the experimental process to ensure reproducibility and reliability of results. Technical replicates and appropriate controls were included in experimental designs to validate findings and minimize experimental variability.
Results:
N-glycosylation: All variants exhibited significant differences in N-glycan structures compared to the wild-type, with increased oligomannose content observed across VOCs. Site-specific analysis revealed alterations in glycan occupancy and composition, indicating variations in glycan processing.
O-glycosylation: Despite being minor, O-glycosylation was present in all variants, predominantly consisting of core-1 O-glycans. Site T323 exhibited conserved O-glycosylation across all variants, with differences in glycan occupancy levels.
Changes in Oligomannose Structures: Variants with higher oligomannose content, particularly Delta and Omicron, showed increased resistance to antibodies, suggesting a potential role in immune evasion.
Conservation of Glycosylation Sites: N-glycosylation sites were highly conserved across variants, highlighting their importance in viral survival and transmission.
Significance of Glycosylation Changes: Glycosylation alterations correlated with increased transmissibility and immune evasion observed in VOCs, emphasizing the role of glycosylation in viral pathogenesis and immune evasion.
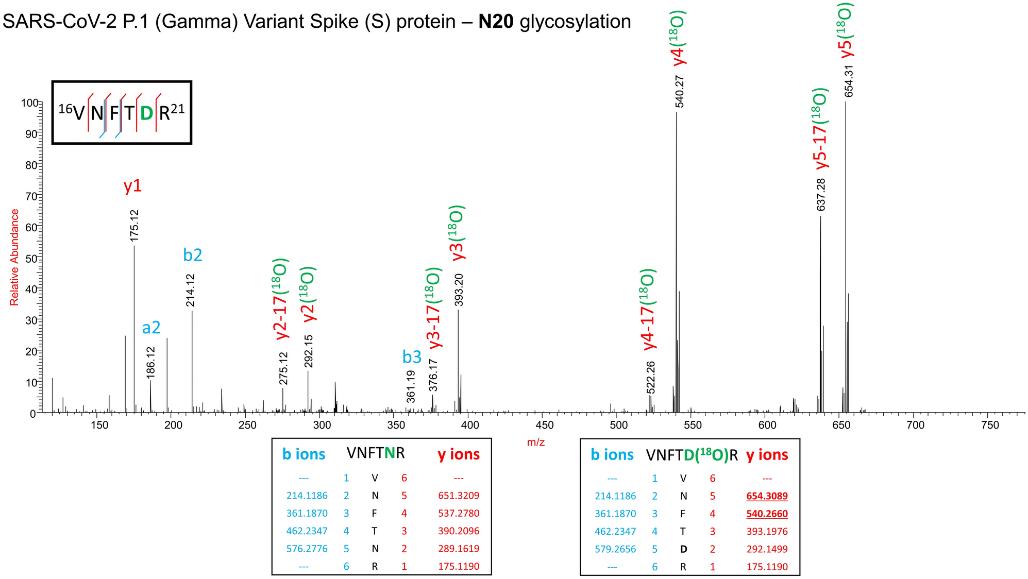 Confirmation of the presence of N-glycosylation at N20 site and absence of N-glycosylation at N17 site on the spike protein of the Gamma variant by stable 18O isotope labelling technique.
Confirmation of the presence of N-glycosylation at N20 site and absence of N-glycosylation at N17 site on the spike protein of the Gamma variant by stable 18O isotope labelling technique.
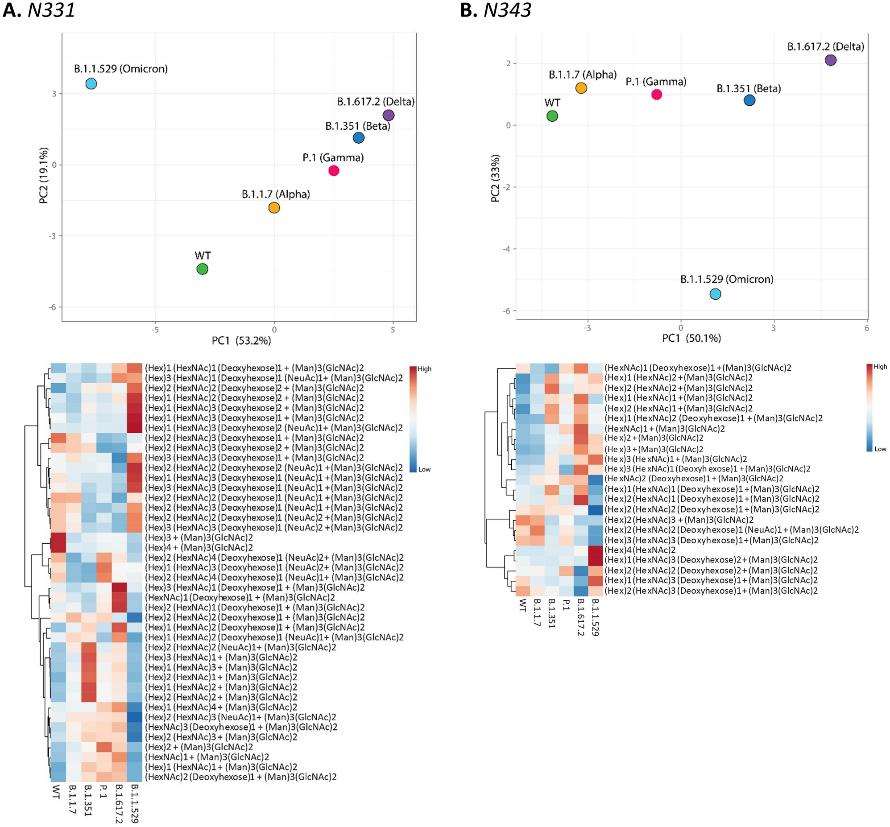 PCA analysis and heat map distribution of the N-linked glycans on sites at RBD, (A) N331 and (B) N343 showed distinct differences in the glycosylation on the VOCs in comparison to the WT, particularly the Omicron variant.
PCA analysis and heat map distribution of the N-linked glycans on sites at RBD, (A) N331 and (B) N343 showed distinct differences in the glycosylation on the VOCs in comparison to the WT, particularly the Omicron variant.
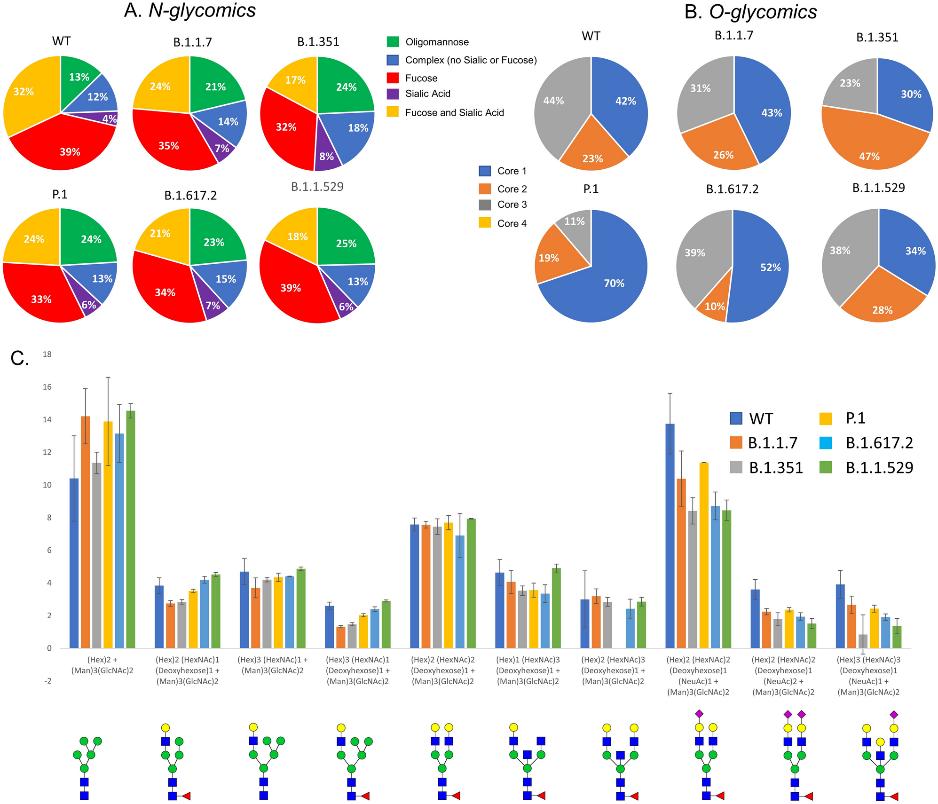 Distribution of (A) N- and (B) O- glycans among the VOCs determined by glycomics (released glycans). All variants showed differences in the type of N-glycans with respect to WT. (C). Relative abundance comparison of most abundant ten glycoforms of WT and VOCs.
Distribution of (A) N- and (B) O- glycans among the VOCs determined by glycomics (released glycans). All variants showed differences in the type of N-glycans with respect to WT. (C). Relative abundance comparison of most abundant ten glycoforms of WT and VOCs.
Reference
- Shajahan, Asif, et al. "Site Specific N-and O-glycosylation mapping of the Spike Proteins of SARS-CoV-2 Variants of Concern." Scientific Reports 13.1 (2023): 10053.




