- Service Details
- Case Study
Introduce of Glycomics
Glycomics aims to decipher the structure, function, and interactions of glycans within the intricate network of biomolecules that comprise living systems. Glycans are not only essential components of cell membranes and extracellular matrices, but they also play crucial roles in numerous physiological processes, including cell-cell recognition, signaling, immune response modulation, and pathogen-host interactions.
The glycome, often referred to as the "sugar code" of life, encompasses the entire repertoire of glycans present in a given biological system under specific conditions of time, space, and environment. This vast and complex landscape of sugars exhibits remarkable diversity in terms of composition, structure, and modification, offering a rich source of information for deciphering biological complexity.
Glycomics employs a wide array of analytical techniques and tools to unravel the intricacies of the glycome. High-resolution mass spectrometry, lectin and antibody arrays, metabolic labeling, and advanced imaging modalities are among the key technologies utilized to characterize glycans at different levels of complexity, from individual structures to global glycomic profiles.
The applications of glycomics span across various fields, including biomedical research, drug development, biotechnology, food science, and environmental science. By elucidating the roles of glycans in health and disease, glycomics holds immense potential for advancing our understanding of biological systems and facilitating the development of novel therapeutics, diagnostics, and biomarkers.
Glycomics Service at Creative Proteomics
Glycan Release Service:
- N-Glycan Liberation Service: Utilizes PNGase F enzyme to selectively release N-glycans from glycoproteins, enabling downstream analysis of N-linked glycosylation patterns.
- O-Glycan Liberation Service: Employ chemical beta-elimination methods, facilitating the release of O-linked glycans for subsequent structural analysis.
Glycan Separation and Purification Services:
Utilize advanced chromatographic techniques to isolate and purify glycan samples, ensuring high-quality preparations for downstream analysis.
Glycan Derivatives Service:
Facilitate glycan modification and functionalization, allowing for enhanced detection, characterization, and manipulation of glycan structures.
Glycan Profiling:
- N-Glycan Profiling: Enable comprehensive analysis of N-linked glycan compositions and structures, providing insights into glycan diversity and heterogeneity.
- O-Glycan Profiling: Offer in-depth characterization of O-linked glycan populations, elucidating their roles in biological systems.
Glycan Structure Analysis:
- Monosaccharide Composition Analysis: Determine the composition of individual monosaccharides within glycan structures, including neutral and amino sugars.
- Oligosaccharide Population Analysis: Allow for the quantification and characterization of glycan oligomers, providing insights into glycan size and complexity.
- Glycan Sequencing: Elucidate the precise sequence of glycan residues, uncovering the linear arrangement of monosaccharides within glycan structures.
- Glycan Linkage Analysis: Enable detailed investigation of glycan linkages, including position, type, branching, and cross-linking analysis.
- Glycan Modification Analysis: Identify and quantify various glycan modifications, including sulfation, acetylation, degradation, and methylation.
Quantitative Glycomics:
- Total Sugar Quantitative Analysis: Provide quantitative measurement of total sugar content within glycan samples, allowing for accurate assessment of glycan abundance.
- Single Glycan Quantitative Analysis: Precise quantification of individual glycan species, enabling comparative studies and biomarker discovery.
Glycomics Data Analysis Services:
Advanced bioinformatics and statistical analysis to interpret glycomic data, providing meaningful insights into glycan structures, compositions, and functions.
Technical Platform for Glycomics Service
At Creative Proteomics, we leverage several technologies and advanced analytical platforms to offer comprehensive glycomics services.
Mass Spectrometry (MS):
Mass spectrometry is a cornerstone technique in glycomics for the analysis of glycan structures. High-resolution MS allows for the accurate determination of molecular weights and fragmentation patterns, facilitating the identification and characterization of individual glycans. Techniques such as MALDI-TOF, ESI-MS, and LC-MS are commonly used to analyze glycan samples, providing valuable information on composition, linkage, and modification.
Lectin and Antibody Arrays:
Lectins, carbohydrate-binding proteins derived from plants and microorganisms, are invaluable tools for glycomic analysis. Lectin arrays enable high-throughput screening of glycan-binding specificities, allowing we to profile the glycome of biological samples rapidly. Similarly, glycan-specific antibodies can be immobilized on arrays to probe glycan expression patterns in a variety of contexts, from cell surfaces to tissue sections.
Metabolic Labeling:
Metabolic labeling techniques involve the incorporation of modified sugar precursors into cellular glycans, enabling the visualization and tracking of glycan biosynthesis in real-time. By introducing functionalized monosaccharides, such as azide or alkyne derivatives, we can selectively label glycans for subsequent detection via bioorthogonal chemistry reactions. This approach provides insights into glycan dynamics and trafficking within living systems.
Advanced Imaging Modalities:
Imaging technologies, such as fluorescence microscopy and mass spectrometry imaging (MSI), offer spatially resolved visualization of glycans within biological samples. Fluorescence microscopy coupled with lectin or antibody staining allows for the localization of specific glycan epitopes in cells and tissues. MSI techniques, on the other hand, provide molecular maps of glycan distributions in biological specimens, enabling the visualization of glycan heterogeneity and spatial organization.
Glycoproteomics:
Glycoproteomics integrates proteomic and glycomic approaches to study glycoproteins, which are proteins modified with glycans. By combining techniques such as affinity enrichment, enzymatic digestion, and mass spectrometry, we can identify glycosylation sites, characterize glycan structures, and quantify glycoprotein abundance in complex biological samples. This comprehensive analysis sheds light on the role of glycoproteins in cellular processes and disease states.
Bioinformatics and Data Analysis:
With the wealth of data generated from glycomic experiments, bioinformatics tools play a crucial role in data processing, analysis, and interpretation. Computational algorithms and databases aid in glycan structure elucidation, glycan identification, and comparative glycomic analysis. Additionally, machine learning approaches are increasingly employed to extract meaningful insights from large-scale glycomic datasets, facilitating the discovery of glycan biomarkers and therapeutic targets.
Sample Requirements for Glycomics Service
| Sample Type | Sample Size Recommendation |
|---|---|
| Serum/Plasma | 25 µl |
| Tissue Biopsy | 5-10 mg |
| Cell Culture | 1-2 x 106 cells |
| Bacterial Culture | 1-5 x 108 CFU |
| Food Product | 100 mg |
| Glycoproteins | 50-100 µg |
Case. Interrogating the Pig Lung Glycome: Insights into Influenza Virus Binding and Host-Pathogen Interactions
Background:
Understanding the interaction between hosts and pathogens is crucial for managing respiratory diseases. The study focuses on elucidating the pig lung glycome to shed light on respiratory illnesses in swine and potentially in humans, particularly regarding influenza virus binding.
Samples:
The research analyzes pig lung tissue obtained from various sources, including commercially available samples and those from germ-free animals. Ethical guidelines are followed for tissue collection and experimentation.
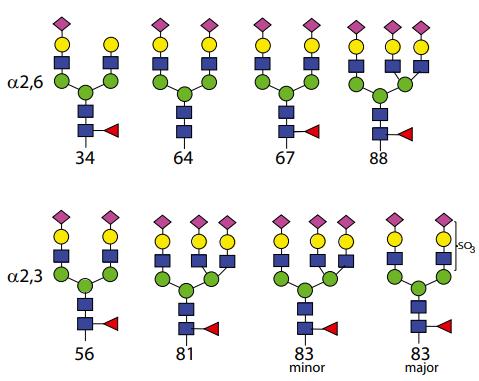 The structures of selected N-glycans.
The structures of selected N-glycans.
Technical Method:
Glycomic Profiling: The pig lung glycome is analyzed using mass spectrometry (MS) to directly examine glycan structures in native tissue. This approach differs from traditional glycan microarray analyses, providing more accurate insights into tissue-specific glycomes.
Glycan Isolation: N-glycans are isolated from pig lung tissue through trypsin digestion and subsequent digestion with Peptide N-Glycosidase F (PNGase F). O-glycans and glycans from glycolipids are isolated using specific procedures. The free reducing glycans are derivatized with 2-aminobenzamide (AEAB) for quantification and microarray analysis.
HPLC Separation: AEAB-conjugated glycans are separated using two-dimensional high-performance liquid chromatography (HPLC) to obtain homogeneous fractions for further analysis.
Virus Preparation: Various influenza virus strains representing swine, human, and avian isolates are grown in Madin-Darby canine kidney (MDCK) cells and purified for subsequent binding assays.
Microarray Preparation: Glycan microarrays are prepared on NHS-activated slides using non-contact printing. The arrays consist of AEAB-conjugated glycans derived from pig lung tissue, allowing for the assessment of virus binding specificity.
Results:
Novel Insights into Glycan Receptor Specificity: The study challenges assumptions about influenza virus binding by revealing unexpected binding patterns and dual specificity for α2,3-Sia and α2,6-Sia glycans among different virus strains.
Characterization of Principal Glycan Binders: Specific glycans containing core fucose and terminal sialic acid are identified as principal binders for multiple virus strains, providing valuable insights into virus-host interactions.
Implications for Host Range and Pathogenicity: Similarities in glycans recognized by swine and human influenza virus strains suggest the importance of glycan receptors in determining host range. Further research into natural receptors is warranted for a comprehensive understanding of virus-host interactions.
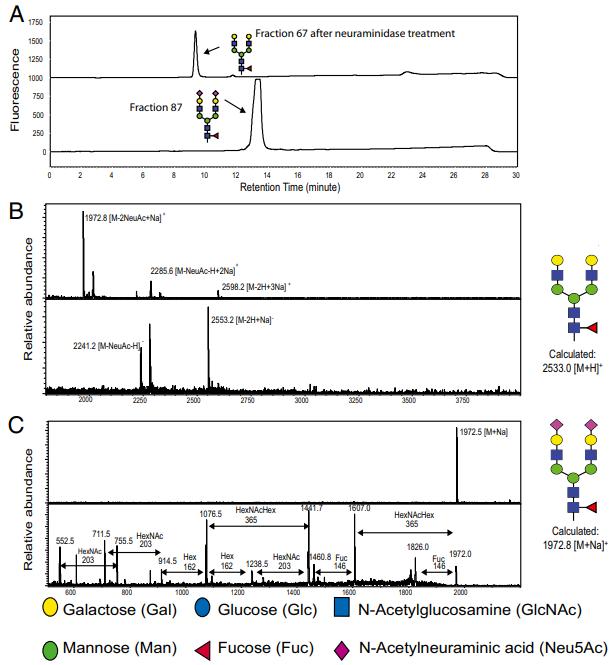 Determination of the structure of fraction 67. (A) HPLC profile of fraction 67 before and after treatment of neuraminidase. (B) The MS spectra of fraction 67, in both positive (Upper) and negative (Lower) mode. (C) The MS (Upper) analysis of fraction 67 after treatment of neuraminidase and MS/MS (Lower) analysis of parent ion 1972
Determination of the structure of fraction 67. (A) HPLC profile of fraction 67 before and after treatment of neuraminidase. (B) The MS spectra of fraction 67, in both positive (Upper) and negative (Lower) mode. (C) The MS (Upper) analysis of fraction 67 after treatment of neuraminidase and MS/MS (Lower) analysis of parent ion 1972
Reference
- Byrd-Leotis, Lauren, et al. "Shotgun glycomics of pig lung identifies natural endogenous receptors for influenza viruses." Proceedings of the National Academy of Sciences 111.22 (2014): E2241-E2250.




