- Service Details
- Case Study
What is Glycoproteomics?
Glycoproteomics is a specialized field within the broader domain of proteomics, focusing specifically on the comprehensive analysis of glycoproteins. Glycoproteins are proteins that have undergone post-translational modification through the attachment of carbohydrate molecules, known as glycans. These glycans are covalently linked to specific amino acid residues within the protein backbone, forming glycoprotein structures with diverse and complex functionalities.
The study of glycoproteomics encompasses a multidisciplinary approach, integrating principles from biochemistry, molecular biology, analytical chemistry, and bioinformatics. Its primary objective is to characterize the structural composition, abundance, localization, and functional significance of glycoproteins within biological systems.
Key aspects of glycoproteomics research include:
Identification and Characterization: Glycoproteomics involves the identification and characterization of glycoproteins present in biological samples. This process typically utilizes high-throughput analytical techniques such as mass spectrometry, which can accurately determine the amino acid sequence of proteins and identify glycan structures attached to specific sites.
Glycan Analysis: A significant focus of glycoproteomics is the analysis of glycans attached to proteins. This entails determining the composition, structure, and heterogeneity of glycans, including their linkage patterns, branching, and modifications. Techniques such as liquid chromatography and lectin affinity chromatography are commonly employed for glycan analysis.
Functional Insights: Understanding the functional roles of glycoproteins is a central objective of glycoproteomics. Glycans play critical roles in various biological processes, including cell-cell communication, immune recognition, protein folding, and pathogen-host interactions. By elucidating the functions of specific glycoproteins, researchers can gain insights into fundamental biological mechanisms and disease pathways.
Disease Biomarker Discovery: Alterations in glycoprotein expression and glycosylation patterns have been associated with numerous diseases, including cancer, autoimmune disorders, and infectious diseases. Glycoproteomics research aims to identify disease-specific glycoprotein biomarkers that can be used for early diagnosis, prognosis, and therapeutic monitoring.
Therapeutic Targeting: Targeting glycoproteins for therapeutic intervention represents a promising strategy in drug discovery and development. By elucidating the molecular mechanisms underlying disease-associated glycoproteins, researchers can identify potential targets for the development of novel therapeutics, including antibodies, small molecules, and glycan-targeting agents.
Glycoproteomics Services Offered by Creative Proteomics
Glycoprotein Separation and Purification Service:
Isolation and purification of glycoproteins from complex biological samples to enhance their detection and analysis.
Glycoprotein Peptide Enrichment Service:
Selective enrichment of glycopeptides for improved sensitivity and coverage in Glycoproteomics analysis.
Glycopeptide Mapping:
Detailed characterization of glycosylation sites and glycan structures attached to proteins, facilitating the elucidation of complex glycoprotein profiles.
Glycoprotein Quantification Service:
Accurate quantification of glycoproteins using label-free or stable isotope labeling methods for comparative analysis and biomarker discovery.
Glycan Structure Analysis:
- Intact Glycoproteins Analysis: Structural analysis of intact glycoproteins to determine their overall composition and modifications.
- Glycan Composition Analysis: Identification and quantification of glycans attached to glycoproteins.
- Glycan Linkage Analysis: Characterization of glycan linkages and branching patterns.
- Sialic Acid Analysis: Quantification of sialic acid content in glycoproteins.
N- and O-Linked Glycosylation Mapping:
- N- and O-Glycan Profiling Service: Comprehensive profiling of N- and O-linked glycans.
- N- and O-Glycosylation Site Analysis Services: Localization of glycosylation sites within protein sequences.
- N- and O-Glycan Linkage Analysis: Determination of glycan linkages and connectivity.
- N- and O-Glycosylation Site Occupation: Assessment of glycosylation site occupancy and occupancy levels.
Glycoprotein Amino Acid Sequence Analysis:
Determination of the amino acid sequences of glycoproteins, including identification of glycosylation sites and modifications.
Protein Glycosylation Analysis:
Comprehensive analysis of protein glycosylation patterns, including glycan structures, attachment sites, and variations.
Glycoprotein Secondary Structure Analysis:
Prediction and characterization of the secondary structure of glycoproteins to assess their folding and stability.
Glycosylation-Protein Interaction Analysis Services:
Investigation of the interactions between glycoproteins and their binding partners to elucidate their functional significance.
Glycoprotein Biomarker Analysis:
Discovery and validation of glycoprotein biomarkers for disease diagnosis, prognosis, and therapeutic monitoring.
Technical Platform for Glycoproteomics Service
Sample Preparation
Before analysis, samples undergo meticulous preparation to ensure optimal glycoprotein extraction and digestion. This involves:
- Denaturation and Reduction: Proteins are denatured to unfold their tertiary structures, followed by reduction of disulfide bonds to facilitate subsequent enzymatic digestion.
- Enzymatic Digestion: Proteins are enzymatically digested using specific proteases to generate peptide fragments, including glycopeptides.
- Glycan Hydrolysis: Glycans attached to proteins are hydrolyzed using glycosidase enzymes, releasing the glycopeptides for downstream analysis.
Chromatographic Separation
Our chromatography systems enable efficient separation of complex mixtures of peptides and glycans, enhancing the resolution and coverage of glycoproteomic analysis:
- High-Performance Liquid Chromatography (HPLC): HPLC is employed for the separation of peptides based on their hydrophobicity, facilitating the isolation of glycopeptides from non-glycosylated peptides.
- Nano Liquid Chromatography (nanoLC): nanoLC offers high sensitivity and resolution for the separation of peptides, particularly useful for the analysis of low-abundance glycopeptides in complex samples.
Mass Spectrometry Analysis
Creative Proteomics utilizes state-of-the-art mass spectrometry systems for sensitive and accurate detection of glycoproteins and glycans:
- Matrix-Assisted Laser Desorption/Ionization Time-of-Flight Mass Spectrometry (MALDI-TOF MS): MALDI-TOF MS provides high-resolution mass spectra, suitable for the analysis of intact glycoproteins and glycans.
- Ultra-High-Performance Liquid Chromatography Tandem Mass Spectrometry (UHPLC-MS/MS): UHPLC-MS/MS offers enhanced sensitivity and sequencing capabilities, enabling the identification and characterization of glycopeptides with high throughput.
Bioinformatics Analysis
Our bioinformatics team employs advanced algorithms and software tools for data processing and interpretation:
- Glycan Identification: Bioinformatics tools are utilized to match experimental mass spectra with databases of known glycan structures, facilitating the identification of glycan compositions and structures.
- Glycosylation Site Mapping: Computational algorithms are employed to map glycosylation sites within protein sequences based on MS/MS fragmentation patterns, enabling the localization of glycan attachments.
- Structural Characterization: Molecular modeling and simulation techniques are used to predict the three-dimensional structures of glycoproteins and glycans, providing insights into their functional properties.
Sample Requirements for Glycoproteomics Service
| Sample Type | Sample Quantity |
|---|---|
| Cell Culture | 1 x 107 cells |
| Tissue Biopsy | 10 - 100 mg |
| Blood Plasma/Serum | 100 - 500 μL |
| Urine | 1 - 5 mL |
| Saliva | 0.5 - 2 mL |
| Cerebrospinal Fluid (CSF) | 0.5 - 1 mL |
| Cell Culture Supernatant | 1 - 5 mL |
| Protein Extracts | 50 - 200 μg |
Case. Comprehensive Analysis of Glycosylation in HIV Immunogen Design
Background:
Glycosylation plays a critical role in HIV immunogen design, as it influences protein folding and antigenicity. Understanding the complex glycan shield of HIV-1 is essential for developing effective vaccines targeting broadly neutralizing antibody (bNAb) epitopes.
Samples:
Various recombinant HIV immunogens, including soluble Env trimers and uncleaved gp140 structures, were analyzed using advanced glycomics and glycoproteomics techniques.
Technical Method:
Glycan Profiling: Quantitative profiling of N-glycans was conducted using ultrahigh-performance liquid chromatography (UPLC) coupled with fluorescence detection. This method allowed for the accurate assessment of oligomannose-type glycans, which are indicative of native-like folding.
Mass Spectrometry: Electrospray ionization (ESI) and matrix-assisted laser desorption/ionization (MALDI) mass spectrometry were employed for structural characterization of released glycans. Collision-induced dissociation (CID) and electron-transfer dissociation (ETD) techniques provided detailed information on glycan composition and glycosidic linkages.
Site-Specific Glycosylation Analysis: Protease digestion and glycopeptide analysis facilitated the determination of glycan occupancy and microheterogeneity at individual glycosylation sites. Label-free quantification approaches were utilized to assess relative glycan abundances.
Intact Mass Spectrometry: Metabolically engineered gp120 was analyzed to assess global glycan site occupancy. This method involved measuring the entire glycoprotein to determine the level of glycan occupancy.
Ion Mobility Mass Spectrometry: Ion mobility mass spectrometry enabled the separation of glycan isomers, providing detailed structural information on glycan profiles and glycosylation patterns.
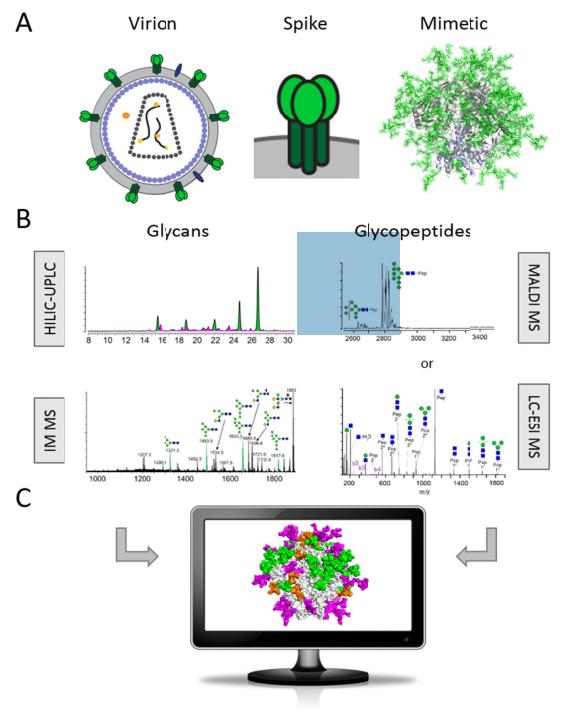 Methods for HIV Env immunogen glycosylation analysis by glycomics and glycoproteomics.
Methods for HIV Env immunogen glycosylation analysis by glycomics and glycoproteomics.
Results:
The comprehensive analysis revealed that leading immunogens of the SOSIP format exhibited native-like folds with a high abundance of oligomannose-type glycans. In contrast, immunogens based on uncleaved gp140 structures showed minimal native-like folding and higher proportions of complex-type glycans. Site-specific glycosylation analysis identified glycan occupancy levels exceeding 90% on recombinant Env trimers. These findings highlighted the importance of glycosylation analytics in evaluating HIV immunogens and understanding bNAb epitopes.
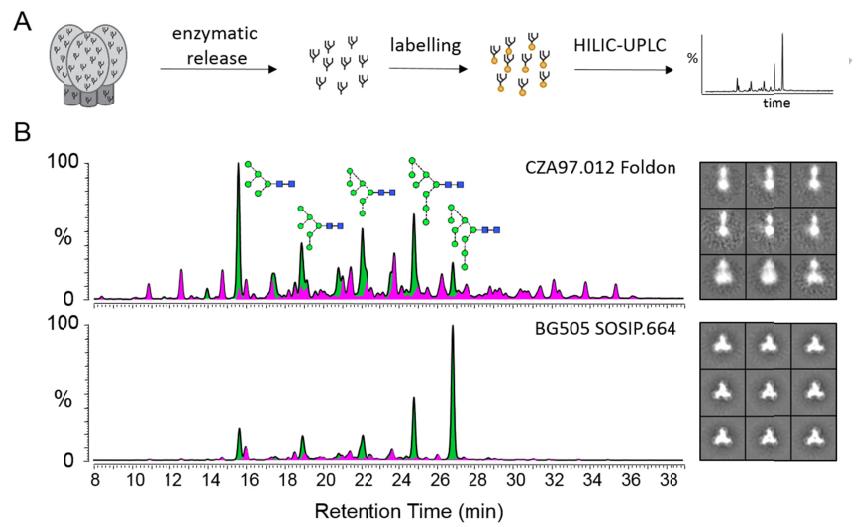 Glycan profiling as a strategy to assess Env immunogen quality.
Glycan profiling as a strategy to assess Env immunogen quality.
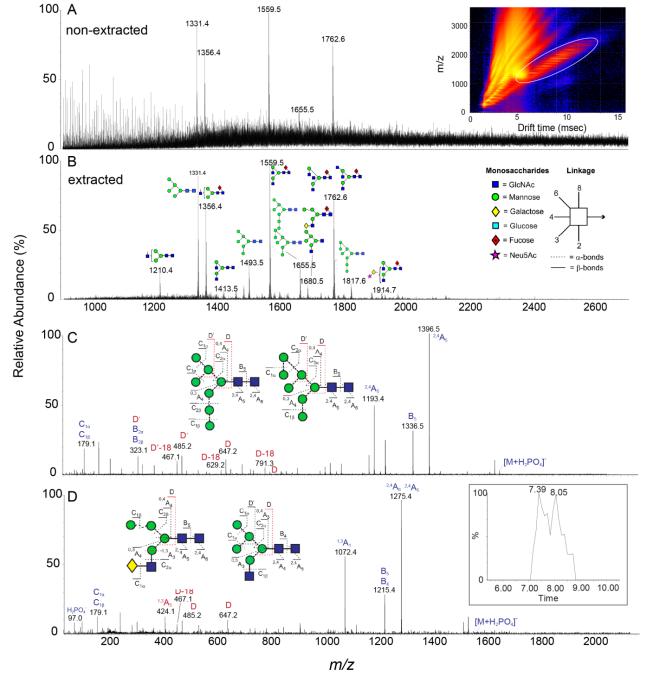 Env glycosylation analysis by ion mobility mass spectrometry.
Env glycosylation analysis by ion mobility mass spectrometry.
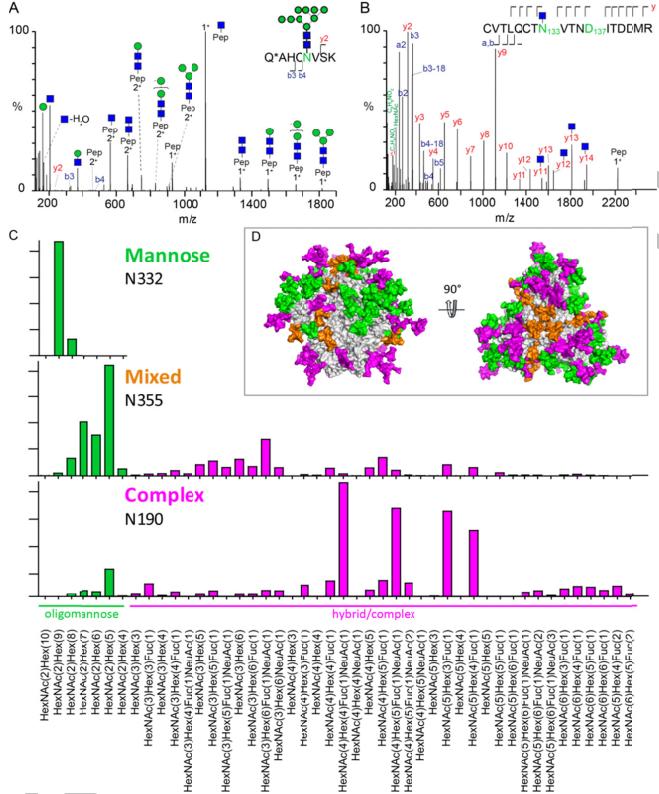 Quantitative site-specific N-glycosylation analysis by LC-ESI MS
Quantitative site-specific N-glycosylation analysis by LC-ESI MS
Reference
- Behrens, Anna-Janina, Weston B. Struwe, and Max Crispin. "Glycosylation profiling to evaluate glycoprotein immunogens against HIV-1." Expert review of proteomics 14.10 (2017): 881-890.




