- Service Details
- Case Study
What is Glycopeptide Mapping?
A deeper understanding of proteins and their functions prompts the need for exploring their molecular architecture. Glycopeptide mapping offers a powerful tool for identifying the complex molecular conformation of glycoproteins, which are proteins adorned with an intricate carbohydrate shell. This technique primarily focusses on understanding post-translational modifications, a burning area of interest in the field of molecular and cellular biology.
The process involves proteolytic digestion of glycoproteins followed by high-resolution mass spectrometry analysis. Combined with innovative technology platforms, this method provides detailed structural information including the identity, location and structural composition of individual glycans on the protein. Thus, glycopeptide mapping serves as an exceptionally important tool in the world of biotechnology, pharmaceutical development, and disease diagnosis.
Technology Platform for Glycopeptide Mapping
At Creative Proteomics, our glycopeptide mapping services are built upon a robust and sophisticated technology platform that integrates cutting-edge liquid chromatography and mass spectrometry techniques.
Ultra-Performance Liquid Chromatography (UPLC)
Our glycopeptide mapping workflow begins with UPLC, a state-of-the-art chromatographic technique known for its exceptional speed, resolution, and sensitivity. UPLC allows for efficient separation of complex peptide mixtures, including glycopeptides, based on their physicochemical properties such as hydrophobicity, size, and charge. With precise control over gradient profiles and column temperatures, we optimize chromatographic conditions to achieve optimal separation and resolution of glycosylated peptides.
High-Resolution Mass Spectrometry (MS)
Coupled with UPLC, our glycopeptide mapping platform incorporates high-resolution mass spectrometry (MS) for accurate and sensitive detection of glycopeptides. High-resolution MS enables precise determination of molecular masses and elucidation of glycan structures attached to specific peptide sequences. By leveraging advanced MS techniques such as electrospray ionization (ESI) and collision-induced dissociation (CID), we achieve comprehensive characterization of glycosylation patterns with unparalleled accuracy and depth of coverage.
Hydrophilic Interaction Liquid Chromatography (HILIC)
A key component of our technology platform is Hydrophilic Interaction Liquid Chromatography (HILIC), a specialized chromatographic mode optimized for the separation of hydrophilic compounds such as glycopeptides. HILIC facilitates selective retention and separation of glycosylated peptides based on their glycan composition, polarity, and size. By employing HILIC columns with tailored stationary phases and mobile phase compositions, we enhance the resolution and specificity of glycopeptide analysis, enabling precise identification and quantification of glycan moieties.
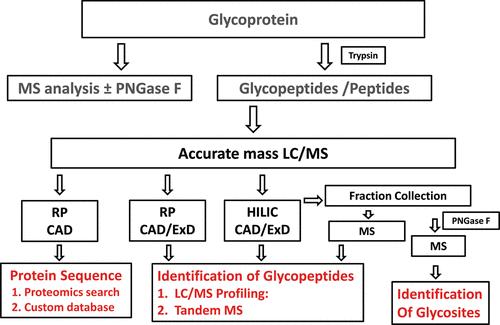 Workflow for N-linked glycopeptide mapping (Leymarie et al., 2012).
Workflow for N-linked glycopeptide mapping (Leymarie et al., 2012).
Data Analysis and Interpretation
Central to our glycopeptide mapping platform is robust data analysis and interpretation methodologies. We utilize sophisticated bioinformatics tools and software algorithms to process and analyze mass spectrometry data, including deconvolution of complex spectra, glycan identification, and peptide mapping. Our team of experienced bioinformaticians and scientists employs advanced statistical methods and database searching algorithms to extract meaningful insights from large-scale glycopeptide datasets, facilitating the elucidation of glycosylation patterns and site-specific heterogeneity.
Quality Control and Assurance
Throughout the glycopeptide mapping process, stringent quality control measures are implemented to ensure the reliability and reproducibility of results. We adhere to established quality assurance protocols and standards, including instrument calibration, method validation, and sample QC/QA procedures. By maintaining rigorous quality control at every step of the analysis workflow, we uphold the highest standards of data integrity and accuracy, providing our clients with confidence in the reliability of our glycopeptide mapping services.
Sample Requirements for Glycopeptide Mapping
| Sample Type | Sample Volume | Additional Requirements |
|---|---|---|
| Monoclonal Antibody | 100 μg (≈ 2-5 μL) | High purity (>95%) |
| Recombinant Protein | 50-100 μg | High concentration (≥1 mg/mL) |
| Serum or Plasma | 100-200 μL | Depletion of high-abundance proteins recommended |
| Cell Culture Medium | 1-2 mL | Filtration or precipitation of proteins required |
| Tissue Samples | 10-50 mg | Homogenization or extraction required |
| Cell Lysates | 1-5 x 106 cells | Lysis buffer optimization required |
| Biopharmaceutical Formulations | 50-100 μg | Sample dilution or buffer exchange may be necessary |
| Body Fluids | 100-500 μL | Sample pre-treatment (e.g., filtration) may be needed |
Applications of Glycopeptide Mapping
Biopharmaceutical Development: Facilitating the characterization of therapeutic glycoproteins, including monoclonal antibodies and recombinant proteins, to ensure product quality and efficacy.
Biomarker Discovery: Identifying glycosylation biomarkers associated with disease states for diagnostic and prognostic applications.
Comparative Glycoproteomics: Comparing glycosylation profiles between different cell lines, production batches, or therapeutic formulations to assess product consistency and comparability.
Structural Elucidation: Unraveling the structure-function relationships of glycoproteins by mapping glycan modifications at specific sites of protein glycosylation.
Why Choose Us?
- Advanced technology: We leverage cutting-edge technologies like HPLC, UPLC, Orbitrap mass spectrometry to ensure high-resolution and accurate data.
- Comprehensive coverage: From small research projects to large scale industrial applications, our services are versatile and scalable.
- Expert analysis: With our skilled bioinformatics team and seasoned analysts, we provide insightful data interpretation.
- Regulatory compliance: Our services are compliant with the necessary regulatory standards ensuring the integrity of your data.
Case. Characterization of Glycosylation Patterns in VEGFR-IgG Fusion Protein Using LC-MS/MS Analysis
Background:
The development of biosimilar drugs, particularly monoclonal antibodies and Fc fusion proteins, has grown significantly to mitigate healthcare costs associated with biopharmaceuticals. Glycosylation, a critical post-translational modification, influences the pharmacokinetics and efficacy of these biotherapeutics. Understanding the site-specific glycosylation patterns is essential for assessing biosimilarity and ensuring the safety and efficacy of biosimilar candidates.
Samples:
The study focused on the VEGFR-IgG fusion protein, a fusion of vascular endothelial growth factor receptors (VEGFR-1 and VEGFR-2) with the Fc portion of human IgG1. Samples included purified fusion protein obtained from cells and cleaved glycans released from the protein.
Technical Method:
Stable Cell Line Generation: Cells were transfected with the expression vector for VEGFR-IgG fusion protein using a recombination-based transfection protocol. High-expressing clones were selected via puromycin selection.
Stable Expression and Purification: Fed-batch culture of transfected cells yielded the fusion protein, which was purified using protein A affinity chromatography.
Cleaved Glycans Preparation: Glycans were released from the protein using PNGase F enzyme, followed by purification using solid-phase extraction (SPE) cartridge.
MALDI MS: Released glycans were analyzed using matrix-assisted laser desorption/ionization mass spectrometry (MALDI-MS) for glycan composition profiling.
Tryptic Digestion and LC-MS/MS Analysis: Protein samples were digested with trypsin, and resulting peptides were analyzed using liquid chromatography-tandem mass spectrometry (LC-MS/MS) for glycopeptide mapping.
N-Glycopeptide Identification: Site-specific glycopeptides were identified using an Integrated GlycoProteome Analyzer (I-GPA), combining glycan-specific oxonium ions and theoretical glycan databases.
Results:
MALDI MS Analysis: Detected major glycan peaks but lacked site-specific information due to glycan pooling.
LC-MS/MS Analysis: Identified N-linked glycosylation sites and characterized glycosylation microheterogeneity.
Site-Specific Glycosylation Patterns: VEGFR-1 showed high fucosylation and sialylation ratios, while VEGFR-2 displayed lower fucosylation and significant sialylation. The Fc region exhibited typical fucosylation patterns observed in human IgG.
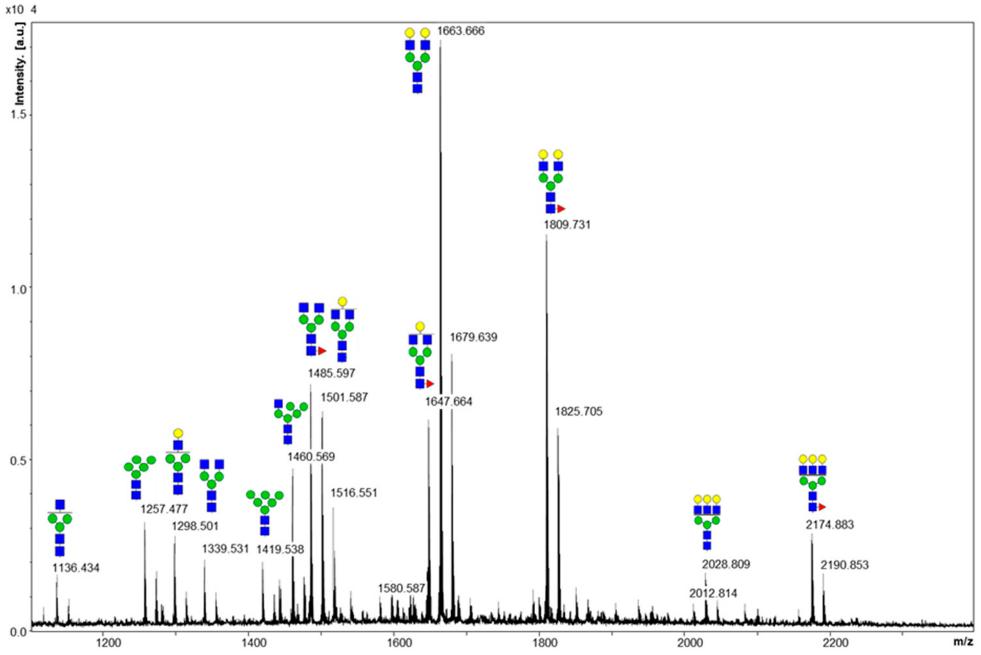 MALDI MS spectrum of the glycans released from the fusion protein. Sodium adducts of each glycan are depicted on the spectrum.
MALDI MS spectrum of the glycans released from the fusion protein. Sodium adducts of each glycan are depicted on the spectrum.
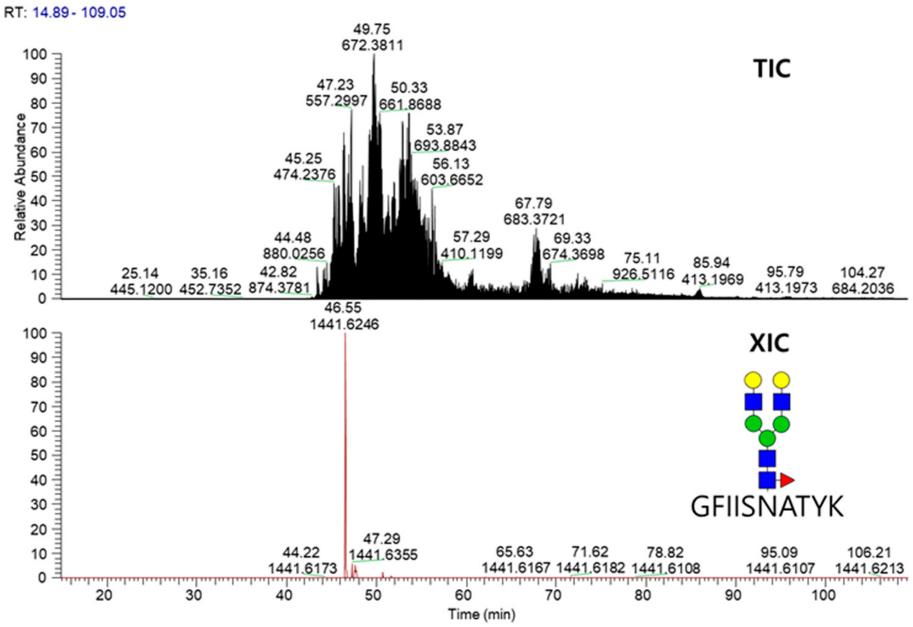 TIC and XIC chromatograms for glycopeptide GFIISNATYK_5_4_1_0 released from the fusion protein.
TIC and XIC chromatograms for glycopeptide GFIISNATYK_5_4_1_0 released from the fusion protein.
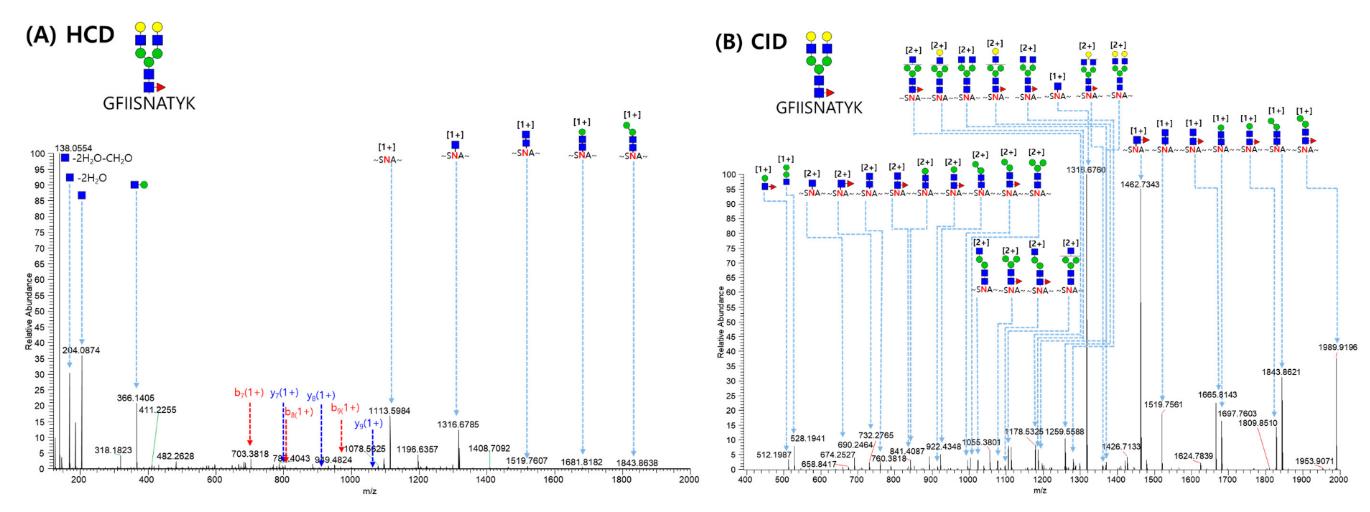 MS/MS spectrum of the GFIISNATYK_5_4_1_0 glycopeptide from the VEGFR-1 region (Asn68) matched automatically by GPA software.
MS/MS spectrum of the GFIISNATYK_5_4_1_0 glycopeptide from the VEGFR-1 region (Asn68) matched automatically by GPA software.
References
- Leymarie, Nancy, and Joseph Zaia. "Effective use of mass spectrometry for glycan and glycopeptide structural analysis." Analytical chemistry 84.7 (2012): 3040-3048.
- Hahm, Young Hye, Ju Yeon Lee, and Yeong Hee Ahn. "Investigation of site-specific differences in glycan microheterogeneity by N-glycopeptide mapping of VEGFR-IgG fusion protein." Molecules 24.21 (2019): 3924.




