- Service Details
- Case Study
What is Glycoprotein Biomarker?
A glycoprotein biomarker, a specialized biological molecule present in blood, bodily fluids, or tissues, undergoes modification through glycosylation, wherein sugar molecules are added to its structure. These biomarkers, complex proteins with glycan modifications, play pivotal roles in various physiological and pathological processes like inflammation, infection, oncogenesis, and tumor progression. Particularly in cancer, alterations in glycosylation status can signify the presence or absence of specific cancer cells, making glycoprotein biomarkers invaluable for disease diagnosis, prognosis, and treatment monitoring.
One distinguishing feature of glycoprotein biomarkers is their capability to mirror changes in an organism's health status. Engaged in cellular activities like signaling, adhesion, and immune response modulation, glycoproteins act as pivotal agents in disease development and progression. Therefore, shifts in their expression levels or glycosylation patterns serve as indicative markers or "biomarkers" for various diseases.
The identification and quantitative analysis of glycoprotein biomarkers offer valuable insights for the development of specific therapeutic strategies in personalized medicine. Through scrutinizing specific glycoproteins' presence, abundance, or modifications in samples like blood, urine, or tissue specimens, researchers gain crucial insights into disease mechanisms. Moreover, identifying and validating novel glycoprotein biomarkers can open avenues for targeted therapeutic interventions and personalized medicine strategies.
Glycoprotein Biomarker Analysis Services in Creative Proteomics
At Creative Proteomics, we take pride in our reputation as a distinguished leader in biomolecular research. Our extensive portfolio includes comprehensive and unique services for glycoprotein biomarker analysis, meticulously designed by seasoned professionals to assist researchers in various applications.
Glycoprotein Isolation and Enrichment
Our glycoprotein isolation and enrichment services are tailored to extract high-purity glycoproteins from complex biological samples or clinical specimens. Leveraging advanced techniques such as lectin-affinity chromatography, hydrazide chemistry, and hydrophilic interaction liquid chromatography (HILIC), we ensure efficient isolation and enrichment of glycoproteins for downstream analysis.
Glycoprotein Identification
Creative Proteomics utilizes mass spectrometry (MS) analysis to rapidly identify glycoproteins in samples. Our glycoprotein identification services enable accurate identification of glycoproteins present, providing valuable insights into their roles in biological processes. Additionally, we offer glycosylation site identification services, employing advanced tandem mass spectrometry to precisely locate glycosylation sites on glycoproteins.
Glycan Profiling and Quantitation
In the final stage of our services, we conduct glycan profiling and glycan quantitation to examine the entire glycome in a biological extract and quantify attached glycans. Our comprehensive approach includes services such as glycan profiling, quantitative glycomics, and glycan structure determination, ensuring a thorough analysis of glycoprotein biomarkers.
N- and O-linked Glycosylation Analysis Service
With the specific N- and O-linked glycosylation analysis services, Creative Proteomics enables researchers to delve into the differential aspects of glycosylation mechanisms.
Technical Platforms for Glycoprotein Biomarker Analysis
Mass Spectrometry (MS): Mass spectrometry-based strategies form the cornerstone of our biomarker discovery workflows. Our diverse range of mass spectrometers, from Matrix-Assisted Laser Desorption/Ionization (MALDI) to Orbitrap, provides unmatched sensitivity and coverage for a wide range of molecular biomarkers. These platforms accommodate various sample types and analytical needs, enabling comprehensive analysis.
High-Resolution Liquid Chromatography (HPLC): Coupled with mass spectrometry, our HPLC systems allow for precise separation and profiling of complex samples. This combination enhances the detection of low-abundance biomarkers that may be overlooked by conventional techniques, providing valuable insights into glycoprotein biomarkers.
Nano-LC-MS/MS Platform: Our highly sensitive and accurate Nano-LC-MS/MS platform is specifically tailored for the analysis of low abundance and complex biological samples. This instrument enables in-depth analysis of glycoprotein biomarkers, contributing to a comprehensive understanding of their roles in health and disease.
Biostatistics: Complementing our experimental platforms is a suite of specialized bioinformatics tools. These tools are designed to process large datasets and extract meaningful insights, facilitating the interpretation of complex data generated from Glycoprotein Biomarker Analysis.
Sample Requirements for Glycoprotein Biomarker Analysis
| Sample Type | Sample Volume Required |
|---|---|
| Serum/plasma | 100-500 μL |
| Urine | 1-5 mL |
| Tissue (e.g., biopsy) | 10-50 mg |
| Cell culture supernatant | 1-5 mL |
| Cerebrospinal fluid | 100-500 μL |
| Saliva | 100-500 μL |
Case. Integrated Glycomic Analysis Reveals Candidate Biomarkers for Hepatocellular Carcinoma Diagnosis
Background:
Hepatocellular carcinoma (HCC) is a leading cause of cancer-related deaths worldwide, and early detection is crucial for effective treatment. Glycomic analysis of serum N-glycans has emerged as a promising approach for identifying biomarkers for HCC diagnosis. However, previous studies have often excluded sialylated glycans, limiting the comprehensive understanding of glycomic alterations in HCC.
Samples:
Serum samples from HCC cases and cirrhotic controls were collected from cohorts in Egypt and the U.S. The cohorts differed in etiological factors, such as hepatitis virus infection status and demographic characteristics.
Technical Method:
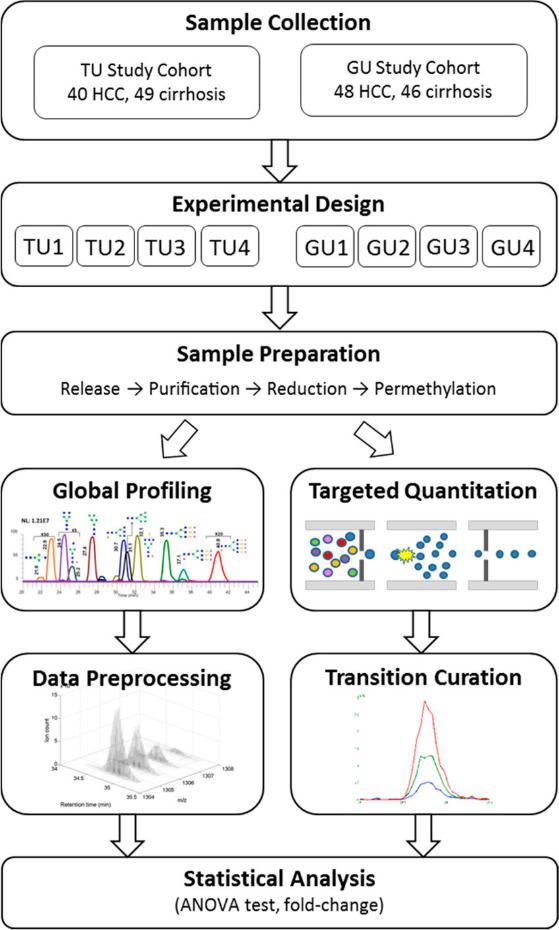
Global Profiling:
- LC–ESI-MS Data Preprocessing: Raw LC–ESI-MS data underwent preprocessing, where peaks outside the expected retention time (RT) range of glycans (15–50 min) were excluded to focus on relevant signals.
- Peak Detection and Batch Matching: Peak detection was performed on the preprocessed data for each batch within the TU and GU cohorts. Detected peaks were then matched across batches within each cohort to ensure consistency.
- Statistical Analysis: A two-way ANOVA model was applied to evaluate the statistical significance of detected peaks. Peaks with a p-value <0.05 and consistent fold changes across the four batches within each cohort were considered significant.
- Glycan Structure Assignment: Putative glycan structures were assigned to the selected peaks by matching experimentally measured mass values with theoretical values of human serum N-glycans, considering different charge states and adduct forms.
Targeted Quantitation:
- Manual Curation of Chromatographic Profiles: Chromatographic profiles were manually curated to eliminate channels with unfavorable profiles or significant noise, ensuring the selection of reliable information for quantitation.
- Multiple Reaction Monitoring (MRM) Analysis: MRM analysis was employed to monitor transitions of N-glycans, allowing for targeted quantitation with improved sensitivity and accuracy compared to global profiling.
- Normalization: A normalization step was applied to ensure consistency across LC–ESI-MRM-MS runs from the same batch, where the mean of log-transformed peak intensities was made identical.
- Statistical Analysis: Similar to global profiling, a two-way ANOVA model was used to identify significant glycans, considering p-values <0.05 and consistent fold changes across batches within each cohort.
Results:
Global Profiling: Identified 26 significant N-glycans, with 18 in the TU cohort and 11 in the GU cohort. Three glycans were significant in both cohorts. Most significant glycans were cohort-specific, reflecting differences in population characteristics and disease stage.
Targeted Quantitation: Detected 15 significant N-glycans, with 11 in the TU cohort and 5 in the GU cohort. Notably, eight glycans overlapped with global profiling results.
Integration and Analysis: A total of 30 unique significant N-glycans were identified, with 11 overlapping between global profiling and targeted quantitation. Clustering analysis revealed patterns in glycan alterations related to biosynthesis processes, such as β-1,6-GlcNAc branching and sialylation.
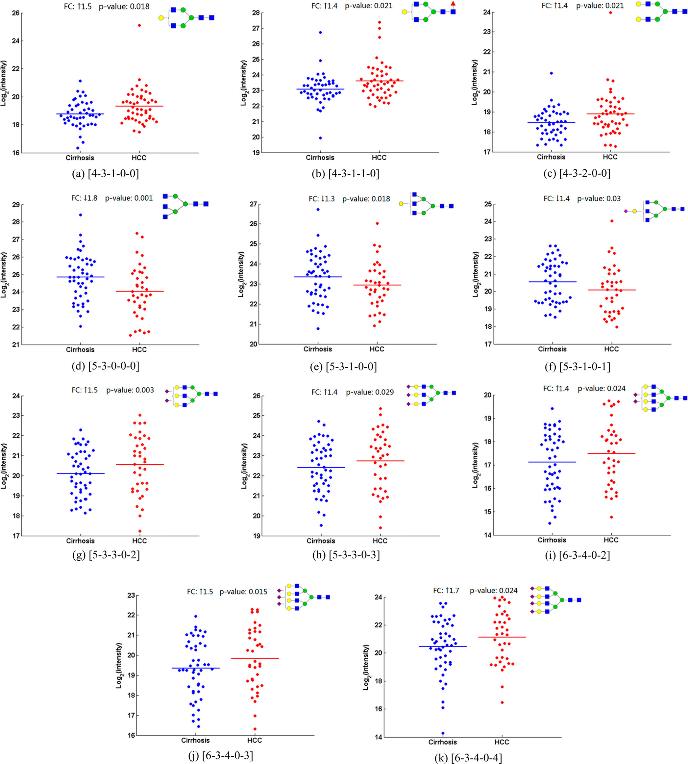 Quantitation results of 11 candidate N-glycan biomarkers in sera of HCC cases and cirrhotic controls by the MRM analysis.
Quantitation results of 11 candidate N-glycan biomarkers in sera of HCC cases and cirrhotic controls by the MRM analysis.
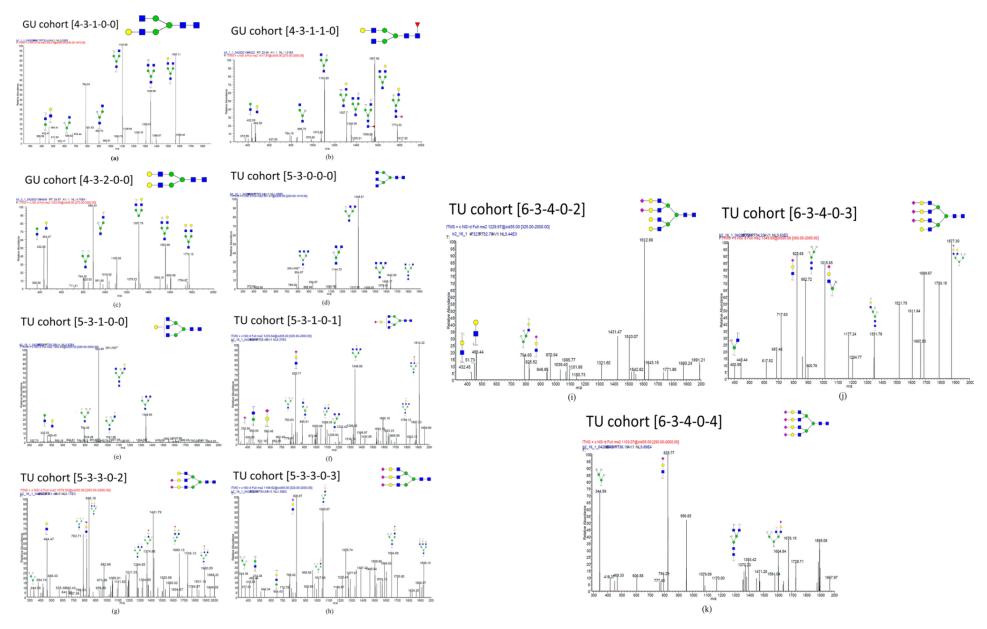 Annotated tandem MS spectra of 11 candidate N-glycan biomarkers: (a) [4-3-1-0-0], (b) [4-3-1-1-0], (c) [4-3-2-0-0], (d) [5-3-0-0-0], (e) [5-3-1-0-0], (f) [5-3-1-0-1], (g) [5-3-3-0-2], (h) [5-3-3-0-3], (i) [6-3-4-0-2], (j) [6-3-4-0-3], and (k) [6-3-4-0-4]. Fragment assignment was based on the criterion of signal-to-noise ratio >3.
Annotated tandem MS spectra of 11 candidate N-glycan biomarkers: (a) [4-3-1-0-0], (b) [4-3-1-1-0], (c) [4-3-2-0-0], (d) [5-3-0-0-0], (e) [5-3-1-0-0], (f) [5-3-1-0-1], (g) [5-3-3-0-2], (h) [5-3-3-0-3], (i) [6-3-4-0-2], (j) [6-3-4-0-3], and (k) [6-3-4-0-4]. Fragment assignment was based on the criterion of signal-to-noise ratio >3.
Reference
- Tsai, Tsung-Heng, et al. "LC–MS profiling of N-glycans derived from human serum samples for biomarker discovery in hepatocellular carcinoma." Journal of proteome research 13.11 (2014): 4859-4868.




