- Service Details
- Case Study
- Description
Peptidoglycan constitutes a fundamental component of bacterial cell walls, characterized by its intricate macromolecular structure composed of diverse amino sugars and peptide chains. Beyond providing structural support and shielding cells from environmental pressures, the bacterial cell wall, wherein peptidoglycan resides, plays pivotal roles in essential physiological processes such as cellular growth, division, and signal transduction. A primary function of peptidoglycan is to uphold the integrity and stability of the cell wall, enabling bacteria to withstand fluctuations in osmotic pressure and exhibit resilience against deleterious agents present in the external milieu.
Analyzing the structure and composition of peptidoglycan is imperative for elucidating the biological characteristics and pathogenic mechanisms of bacteria, as well as understanding the mechanisms of action of antibiotics. Through in-depth investigation into various aspects of peptidoglycan, including its composition, modifications, and cross-linkages, researchers can discern structural disparities among different bacterial strains and identify novel drug targets. Moreover, peptidoglycan analysis aids in the assessment of bacterial resistance to antibiotics, thereby offering critical insights for optimizing antibiotic therapies and facilitating the development of novel therapeutic agents. Consequently, peptidoglycan analysis holds significant promise in both microbial research and clinical therapeutics.
Peptidoglycan Analysis Services in Creative Proteomics
Peptidoglycan Composition Analysis: Identify and quantify the individual components of peptidoglycan, providing insights into structural diversity among bacterial species or strains.
Peptidoglycan Structure Elucidation: Determine the detailed arrangement of sugars, amino acids, modifications, and cross-linkages within peptidoglycan molecules.
Peptidoglycan Modification Analysis: Characterize modifications such as glycosylation or acetylation, uncovering mechanisms of bacterial pathogenesis and drug resistance.
Peptidoglycan Cross-Linking Analysis: Analyze patterns and extent of cross-linking, crucial for understanding cell wall dynamics and developing antibacterial strategies.
Peptidoglycan Turnover Dynamics Study: Investigate kinetics of peptidoglycan synthesis, degradation, and recycling, providing insights into bacterial growth and response to stresses.
Technical Platforms for Peptidoglycan Analysis
High-Performance Liquid Chromatography (HPLC): HPLC systems equipped with UV or fluorescence detectors for precise quantification of peptidoglycan components.
Mass Spectrometry (MS): MALDI-TOF or LC-MS instruments for high-resolution structural analysis, enabling identification of peptide sequences and modifications.
Nuclear Magnetic Resonance (NMR) Spectroscopy: NMR spectrometers for elucidating three-dimensional peptidoglycan structures and dynamics with atomic resolution.
Glycan Microarrays: High-throughput screening platforms to study peptidoglycan interactions with antibodies, proteins, and other ligands.
Fluorescence Microscopy: Fluorescence-labeled probes and confocal microscopes for visualizing peptidoglycan localization and dynamics in living cells.
Enzymatic Digestion and Analysis: Utilization of lytic enzymes coupled with chromatographic or spectrometric techniques for studying peptidoglycan turnover and remodeling dynamics.
Sample Requirements for Peptidoglycan Analysis
| Sample Type | Sample Amount Required |
|---|---|
| Bacterial Cultures | 1-5 mL of culture broth |
| Bacterial Pellets | 1-5 mg of bacterial pellet |
| Bacterial Cell Walls | 1-5 mg of purified cell walls |
| Clinical Samples | Variable, depending on the sample source and method of extraction (e.g., tissue samples, body fluids) |
| Environmental Samples | Variable, depending on the sample source (e.g., soil, water, biofilms) and method of extraction |
| Biofilm Samples | Variable, depending on the size and thickness of the biofilm |
| Synthetically Prepared Peptidoglycan | Variable, depending on the desired concentration and purity of the peptidoglycan |
| Model Organisms or Cell Lines Expressing Modified Peptidoglycan | Variable, depending on the experimental setup and the level of peptidoglycan modification |
| Mutant Bacterial Strains with Altered Peptidoglycan Composition | Variable, depending on the growth conditions and the extent of the peptidoglycan alteration |
Applications of Peptidoglycan Analysis
Antibiotic Development: Essential for creating and enhancing antibiotics by targeting the enzymes involved in peptidoglycan biosynthesis.
Drug Resistance Mechanisms: Helps understand how bacteria develop resistance to antibiotics through changes in peptidoglycan synthesis pathways.
Bacterial Identification: Useful in differentiating and identifying bacterial species based on variations in peptidoglycan structure.
Host-Pathogen Interactions: Critical for studying how peptidoglycan fragments interact with the host immune system, influencing disease outcomes and immune responses.
Diagnostic Biomarkers: Peptidoglycan components can serve as biomarkers for diagnosing bacterial infections.
Vaccine Development: Peptidoglycan components are explored as targets for vaccines designed to elicit protective immune responses against bacterial pathogens.
Title: High-Resolution Analysis of Enzymatically Digested Mycobacterial Cell Walls Using LC-MS
Background
The study aims to analyze the structural components of the mycobacterial cell wall, specifically from Mycobacterium smegmatis (mc2155), which serves as a model for more pathogenic strains like Mycobacterium tuberculosis. Understanding these structures aids in comprehending the bacterial resistance mechanisms and can guide new therapeutic strategies.
Sample
The sample studied is the cell wall of Mycobacterium smegmatis (mc2155), a model organism for mycobacterial research due to its non-pathogenic nature and similarity to pathogenic mycobacteria. The cell walls are isolated from cultures grown to stationary phase in supplemented Middlebrook 7H9 broth.
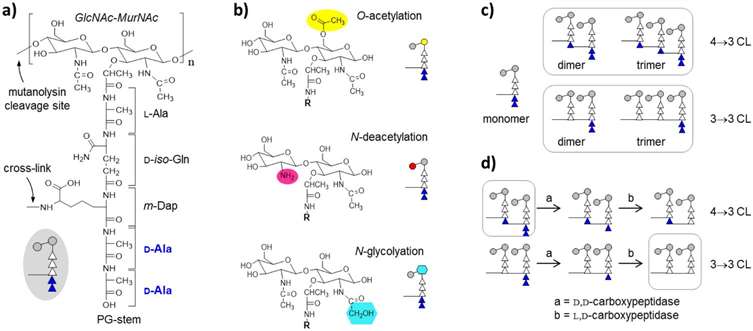 The peptidoglycan of Mycobacterium smegmatis.
The peptidoglycan of Mycobacterium smegmatis.
Technical Methods
- Cultivation and Isolation: M. smegmatis is cultured in Middlebrook 7H9 broth, harvested at stationary phase, and subjected to cell wall isolation involving centrifugation, washing, and SDS treatment.
- Enzymatic Digestion: The isolated cell walls undergo enzymatic digestion using DNase and Pronase E to remove nucleic acids and proteins, followed by mutanolysin to break down peptidoglycan (PG) structures. The digestion processes facilitate the breakdown of complex cell wall components into simpler muropeptide fragments.
- LC-MS Analysis: The digested samples are prepared for LC-MS by reducing with sodium borohydride, quenching with phosphoric acid, filtering, and resuspending in sample preparation buffer. They are then purified using C18 tips and analyzed using a high-resolution Waters C18 nanoACQUITY UPLC coupled with a Synapt G2 HDMS time-of-flight mass spectrometer. The analysis includes specific conditions for elution and gradient settings tailored for optimal separation and detection of muropeptide fragments.
Results
While specific results are not detailed in the provided text, the described methodology implies the generation of detailed mass spectrometric data on muropeptide fragments. This data is crucial for constructing a comprehensive profile of the biochemical structure and modifications within the M. smegmatis cell wall. The study likely results in a high-resolution map of the cell wall's molecular composition, aiding further biological and pharmaceutical research.
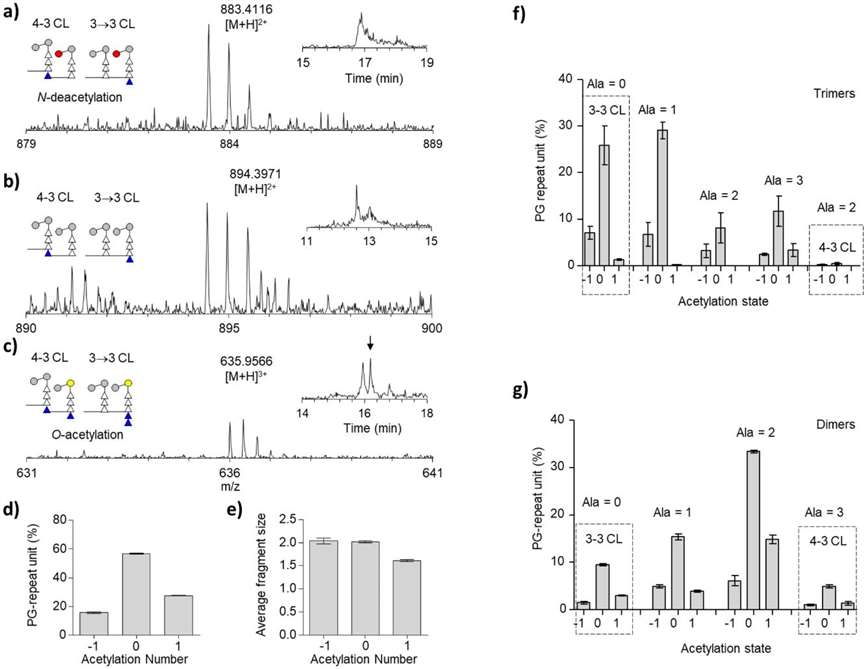 Peptidoglycan acetylation of mc2155 cell walls.
Peptidoglycan acetylation of mc2155 cell walls.
Reference
- Rimal, Binayak, et al. "Peptidoglycan compositional analysis of Mycobacterium smegmatis using high-resolution LC–MS." Scientific reports 12.1 (2022): 11061.
What is peptidoglycan and its function?
Peptidoglycan, also known as murein, is a complex polymer that forms a mesh-like layer outside the plasma membrane of most bacteria. This structure is composed of sugars and amino acids, forming a lattice that provides structural strength and shapes the bacterial cell.
Structure of Peptidoglycan
The backbone of peptidoglycan consists of two alternating sugars: N-acetylglucosamine (NAG) and N-acetylmuramic acid (NAM). These sugars are linked by β-(1,4)-glycosidic bonds. Attached to the NAM sugar is a peptide chain of three to five amino acids, which can cross-link with peptide chains on other strands. This cross-linking provides the structural integrity and rigidity necessary for the survival of bacteria under various conditions.
Functional Role
Peptidoglycan serves several critical functions:
- Structural Support: It maintains the shape of the bacteria and counteracts the internal osmotic pressure of the cell, preventing lysis.
- Barrier: Acts as a protective barrier against environmental threats, including toxins and antibiotics.
- Growth and Division: It is dynamically remodeled during cell growth and division, allowing for the expansion and safe division of the bacterial cell wall.
How is peptidoglycan important in identifying specific bacteria?
The structure of peptidoglycan varies significantly among different bacterial species, making it a valuable phenotypic marker for bacterial identification. The variations in peptidoglycan structure can affect the permeability, rigidity, and overall susceptibility of bacteria to antibiotics, influencing their detection and characterization.
Applications in Bacterial Typing:
- Taxonomic Classification: Differences in peptidoglycan types help classify bacteria into major groups, such as Gram-positive and Gram-negative, based on their cell wall composition.
- Pathogen Identification: Specific modifications in the peptidoglycan layer can signal the presence of pathogenic bacteria, aiding in rapid diagnosis and treatment.
Does Gram stain detect peptidoglycan?
Yes, the Gram stain technique is a fundamental method that indirectly detects peptidoglycan. Developed by Hans Christian Gram, this method exploits the differences in the peptidoglycan layer thickness between Gram-positive and Gram-negative bacteria.
Gram Staining Procedure
- Application of Crystal Violet: The primary stain that penetrates all bacteria.
- Application of Iodine: Forms a complex with the crystal violet that is trapped within the thick peptidoglycan layer of Gram-positive bacteria.
- Decolorization: Alcohol or acetone is used, which dehydrates the peptidoglycan layer, trapping the dye in Gram-positive cells. It washes away the dye from Gram-negative bacteria with thinner peptidoglycan.
- Counterstaining: Typically with safranin, which stains Gram-negative bacteria red or pink.
What stains positive for peptidoglycan?
In the context of the Gram stain, Gram-positive bacteria stain positive for peptidoglycan because of their thick peptidoglycan layer which traps the crystal violet dye. Other stains that are sensitive to peptidoglycan include:
- Antibiotic labeling: Binds specifically to the D-Ala-D-Ala terminals in peptidoglycan.
- Fluorescent Dyes: Some fluorescent dyes are designed to bind to peptidoglycan, providing valuable tools for studying bacterial cell walls in live cells.




