What is Glycosphingolipid Microarray Assay?
A glycosphingolipid microarray assay is a powerful analytical technique that enables the high-throughput screening and profiling of glycosphingolipids—complex lipids that play pivotal roles in cell signaling and membrane dynamics. These assays facilitate the investigation of glycosphingolipid interactions with proteins, antibodies, and cells, making them indispensable tools in both basic and applied research.
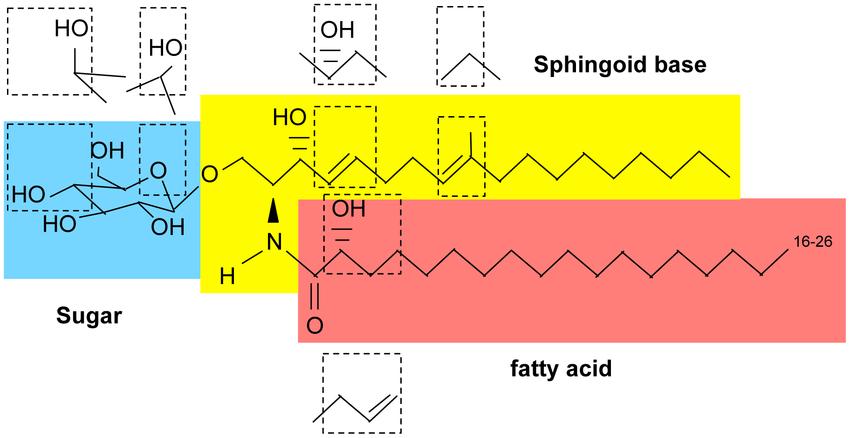 Basic structure of glycosphingolipids (Del Poeta et al., 2015).
Basic structure of glycosphingolipids (Del Poeta et al., 2015).
Principle of Glycosphingolipid Microarray
The principle underlying Glycosphingolipid Microarray Assay revolves around the immobilization of diverse GSLs onto a solid support, typically a glass slide, in a spatially defined array format. This array enables the simultaneous screening of multiple GSLs for their interactions with various biomolecules. The assay facilitates the elucidation of GSL-binding specificities, affinities, and functional consequences, thereby providing invaluable insights into glycosphingolipid-mediated biological processes.
Workflow of Glycosphingolipid Microarray Assay
1) Extraction and isolation of glycosphingolipids (GSLs) from biological samples (tissues, cells, or fluids). This is done by homogenizing the sample in a chloroform/methanol mixture to extract lipids, followed by various purification steps to isolate the GSL fraction.
2) N-monodeacylation of the purified GSLs using an enzyme like sphingolipid ceramide N-deacylase. This exposes an amine group on the GSL for fluorescent labeling.
3) Fluorescent labeling of the N-monodeacylated GSLs at the newly exposed amine group.
4) Printing the fluorescently labeled GSLs onto a microarray slide in a high-density array format, with each GSL printed in replicate spots.
5) Blocking the microarray slide to prevent non-specific binding.
6) Incubating the blocked GSL microarray with the glycan-binding protein (lectin, antibody, etc.) of interest that is fluorescently labeled or will be detected via a secondary labeling step.
7) Washing away any unbound protein to remove non-specific binding.
8) Scanning the microarray slide with a fluorescence scanner to detect and quantify the binding of the protein to each printed GSL glycan.
9) Data analysis to determine the binding profiles and specificities of the glycan-binding protein towards the various GSL glycans present on the array.
Glycosphingolipid Microarray Assay Platform Offered by Creative Proteomics
High-Throughput Screening: Simultaneous analysis of multiple GSL interactions with proteins, antibodies, lectins, and other biomolecules for accelerated research outcomes.
GSL-Binding Specificity Profiling: Comprehensive profiling of GSL-binding specificities to elucidate molecular recognition events and identify potential ligands.
Affinity Determination: Quantitative assessment of GSL-binding affinities towards biomolecules to understand the strength and dynamics of molecular interactions.
Functional Characterization: Investigation of functional consequences of GSL interactions, including signaling pathways modulation, immune response modulation, and cell-cell recognition.
Customized Assay Design: Tailored assay design and optimization to meet specific research objectives and experimental requirements.
Data Analysis and Interpretation: Comprehensive data analysis and interpretation services to extract meaningful insights from GSL microarray results.
Advantages of Our Glycosphingolipid Microarray Platform
- High-throughput screening capability: It allows simultaneous analysis of hundreds of glycosphingolipid glycan structures against a glycan-binding protein of interest in a single experiment. This high-throughput nature enables rapid profiling of binding specificities.
- Low sample requirements: Only minute amounts of the GSL sample and the protein are needed for the microarray analysis. This is particularly useful when working with limited clinical samples.
- Comprehensive profiling: The microarray provides detailed information about the glycan linkages and structural features that govern the binding interactions with proteins. This aids in deciphering the glycan structural basis for biological functions.
- Simple workflow: The microarray assay has a relatively simple workflow involving GSL extraction, labeling, printing on slides, incubation with proteins, and fluorescence detection. This ease of use enables high-throughput screening.
- Cost-effective: Compared to traditional techniques like mass spectrometry, the glycosphingolipid microarray is a more cost-effective platform for initial screening and studying GSL-protein interactions.
- Versatile applications: The microarrays can be used to study interactions with various biomolecules like antibodies, lectins, cells, bacteria, viruses etc., enabling diverse applications in biomarker discovery, immunology, and pathogen interactions.
Sample Requirements for Glycosphingolipid Microarray Assays
| Sample Type | Sample Quantity | Description |
|---|---|---|
| Purified Proteins | 10-50 µg per replicate | Highly purified proteins of interest, such as receptors or enzymes, for probing GSL interactions. |
| Monoclonal Antibodies | 5-20 µg per replicate | Monoclonal antibodies targeting specific epitopes for assessing GSL-binding specificity. |
| Lectins | 1-10 µg per replicate | Lectins with known carbohydrate-binding specificities for studying GSL-lectin interactions. |
| Cell Lysates | 50-100 µg per replicate | Cell lysates containing GSL-binding proteins for identifying endogenous GSL-protein interactions. |
| Tissue Extracts | 100-500 µg per replicate | Tissue extracts enriched in GSLs for examining tissue-specific GSL interactions. |
| Synthetic Compounds | Variable depending on compound | Synthetic compounds or small molecules for evaluating GSL-binding affinity and specificity. |
Applications of Glycosphingolipid Microarray
Glycan-Protein Interactions
- Glycan Binding Specificity: Profiling the binding specificities of proteins, antibodies, lectins, and other biomolecules towards diverse GSL structures.
- Glycan Binding Affinity: Quantifying the affinity of proteins for specific GSLs to understand the strength and dynamics of glycan-protein interactions.
Glycan Function in Biological Systems
- Cellular Signaling: Investigating the role of GSLs in cellular signaling pathways, including cell adhesion, migration, and differentiation.
- Immune Response: Studying GSL-mediated immune responses, such as antigen recognition by T cells and B cells, and modulation of immune cell function.
Disease Biomarkers and Therapeutics
- Cancer Biomarkers: Identifying GSL biomarkers associated with cancer progression, metastasis, and drug resistance for diagnostic and therapeutic purposes.
- Infectious Diseases: Characterizing GSL interactions with pathogens, such as viruses and bacteria, to develop novel therapeutics and vaccines.
Drug Discovery and Development
- Target Identification: Screening for novel GSL-binding targets for drug discovery and development, including small molecule inhibitors and biologics.
- Drug Efficacy Testing: Assessing the efficacy of GSL-targeting drugs in preclinical studies and clinical trials for personalized medicine approaches.
Glycomics and Systems Biology
- Glycomic Profiling: Mapping the glycome landscape of cells, tissues, and organisms to understand the role of GSLs in biological systems.
- Systems Biology Modeling: Integrating GSL microarray data with other omics datasets to generate comprehensive models of cellular processes and disease mechanisms.
Diagnostic Assays and Biomarker Discovery
- Disease Diagnosis: Developing GSL-based diagnostic assays for diseases, such as lysosomal storage disorders, autoimmune diseases, and neurodegenerative disorders.
- Biomarker Discovery: Identifying novel GSL biomarkers associated with disease pathogenesis, prognosis, and treatment response for precision medicine applications.
Reference
- Del Poeta, Maurizio, et al. "Correction: synthesis and biological properties of fungal glucosylceramide." PLoS Pathogens 11.5 (2015): e1004886.




