- Service Details
- Case Study
What is Label-free Quantitative Analysis?
Label-free quantitative analysis has revolutionized biological research, offering a robust and efficient method for analyzing biomolecules without the need for fluorescent tags or radiolabels. This advanced approach leverages mass spectrometry techniques to directly identify and quantify proteins based on their mass-to-charge ratios, ensuring accurate results and eliminating potential discrepancies from labeling.
The advantages of label-free quantitative analysis are manifold. It saves time and costs by eliminating the need for labeling, allows for the analysis of larger protein sets, and offers an extensive dynamic range suitable for complex biological systems. This technology has profound implications for understanding physiological and pathological processes at the protein level, including the profiling of entire proteomes.
In the realm of glycoproteomics, label-free quantitative analysis holds significant promise. By deciphering the intricate interactions of glycoproteins, researchers can glean invaluable insights into various diseases and biological phenomena. At Creative Proteomics, we harness the power of this innovative technology to deepen our understanding of protein functions and drive advancements in drug development, therapies, and diagnostics. Through precise implementation of label-free quantitative analysis, we unlock the full potential of biotechnology to propel scientific discovery and improve human health.
Creative Proteomics offers a range of specialized projects for glycoprotein label-free quantitative analysis, including:
Glycoprotein Profiling: Comprehensive analysis of glycoproteins to elucidate their structural diversity and functional relevance in biological systems.
Differential Expression Analysis: Quantification of changes in glycoprotein abundance under different experimental conditions, providing insights into disease mechanisms and biomarker discovery.
Glycosylation Site Mapping: Identification and characterization of glycosylation sites on proteins, facilitating the understanding of post-translational modifications and protein function.
Functional Annotation: Integration of quantitative proteomic data with functional databases to annotate glycoproteins and unravel their roles in cellular pathways and disease processes.
Customized Projects: Tailored solutions and optimization strategies to address specific research objectives and sample types, ensuring the success of glycoprotein label-free quantitative analysis projects.
Technology Platform for Glycoprotein Label-free Quantitative Analysis
To efficiently serve the evolving scientific community, we employ a two-fold technological format: mass spectrometry (MS) and liquid chromatography (LC). The combination, known as LC-MS, forms the nucleus of our label-free quantitative analysis at Creative Proteomics.
Mass spectrometry, one of the most widely used analytical techniques in proteomics, takes the center stage in our platform. It allows an in-depth examination and identification of glycoproteins, thanks to its high-resolution power. MS achieves this by measured the mass-to-charge ratio of charged particles, providing us with a spectrum that acts as a unique identifier for each glycoprotein.
While mass spectrometry provides a robust backbone, liquid chromatography acts as a complementary step necessary for label-free quantitative analysis of glycoproteins. LC helps in the separation of complex glycoprotein mixtures based on their physicochemical properties, streamlining the MS analysis. Through the high-pressure application, LC assists in a more precise separation that reduces sample complexity, enabling a more accurate label-free quantification, thus illustrating the synergy between LC and MS at Creative Proteomics.
Sample Requirements for Glycoprotein Label-free Quantitative Analysis
| Sample Type | Sample Volume | Sample Preparation |
|---|---|---|
| Cell Culture | 1-2 x 107 cells | Harvest cells and extract proteins using lysis buffer. |
| Tissue | 10-50 mg | Homogenize tissue in lysis buffer to extract proteins. |
| Serum/Plasma | 100-500 µL | Precipitate proteins with organic solvent or TCA/acetone. |
| Urine | 1-5 mL | Centrifuge urine to remove debris before analysis. |
| CSF (Cerebrospinal Fluid) | 100-500 µL | Centrifuge CSF to remove cells and debris. |
| Saliva | 500 µL - 1 mL | Centrifuge saliva to remove cells and debris. |
Applications of Glycoprotein Label-free Quantitative Analysis
Biomarker Discovery and Validation
Identification of glycoprotein biomarkers associated with disease states, facilitating early diagnosis, prognosis, and monitoring of diseases such as cancer, cardiovascular disorders, and neurological conditions.
Validation of candidate biomarkers through quantitative analysis of glycoprotein expression levels in biological samples from patient cohorts and healthy controls.
Pharmacological Studies and Drug Development
Evaluation of drug-induced changes in glycoprotein expression profiles and post-translational modifications to assess drug efficacy, toxicity, and mechanism of action.
Screening and characterization of glycoprotein targets for drug development and therapeutic interventions, including monoclonal antibodies, vaccines, and small molecule inhibitors.
Functional Characterization of Glycoproteins
Elucidation of the structural diversity and glycosylation patterns of glycoproteins to decipher their roles in cell signaling, immune response, cell adhesion, and protein-protein interactions.
Mapping of glycosylation sites and determination of glycan structures on specific glycoproteins to understand their functional implications in disease progression and physiological processes.
Systems Biology and Pathway Analysis
Integration of glycoprotein quantitative data with other omics datasets (e.g., genomics, transcriptomics, metabolomics) to reconstruct signaling pathways, regulatory networks, and cellular processes underlying complex diseases.
Profiling of entire glycoproteomes in different cell types, tissues, and biological fluids to gain insights into the global changes in glycoprotein expression and glycosylation associated with physiological and pathological conditions.
Vaccine Development
Investigation of glycoprotein antigens for vaccine development against infectious diseases, including viruses, bacteria, and parasites.
Quantitative analysis of glycan structures and glycosylation patterns on vaccine candidates to optimize antigenicity, immunogenicity, and vaccine efficacy.
Environmental Monitoring and Bioprocessing
Monitoring of glycoprotein biomarkers in environmental samples, such as water, soil, and air, for assessing environmental contamination, exposure to pollutants, and ecological health.
Optimization of bioprocessing conditions in biopharmaceutical manufacturing and biotechnological applications through real-time monitoring of glycoprotein expression levels and glycosylation patterns.
Case. Comprehensive Urinary Glycoprotein Profiling for Bladder Cancer Detection
Background:
Bladder cancer diagnosis traditionally relies on invasive procedures like cystoscopy. Urinary proteomics offers a noninvasive approach for biomarker discovery, but existing methods have limitations. This study aims to address these limitations by employing liquid chromatography/tandem mass spectrometry (LC/MS-MS) for comprehensive urinary glycoprotein profiling.
Samples:
The study analyzed urine samples from 54 bladder cancer patients and 46 noncancer controls. The samples were enriched for glycoproteins using dual lectins (ConA and WGA) and subjected to trypsin digestion before LC/MS-MS analysis.
Technical Method:
Enrichment of glycoproteins: Urine samples were passed through columns packed with agarose-bound ConA and WGA lectins.
Tryptic digestion: The enriched glycoproteins were digested with trypsin to generate peptide mixtures.
LC/MS-MS analysis: Peptide mixtures were analyzed using nano-reverse phase liquid chromatography coupled with electrospray ionization mass spectrometry (LTQ).
Database queries and manual validation: MS/MS spectra were analyzed using the SEQUEST algorithm and validated against the Swiss-Prot database.
Differential expression analysis: Spectral counting quantification was used to identify differentially expressed glycoproteins between cancer and noncancer samples.
Validation: Western blotting and ELISA were employed to validate the differential expression of target glycoproteins.
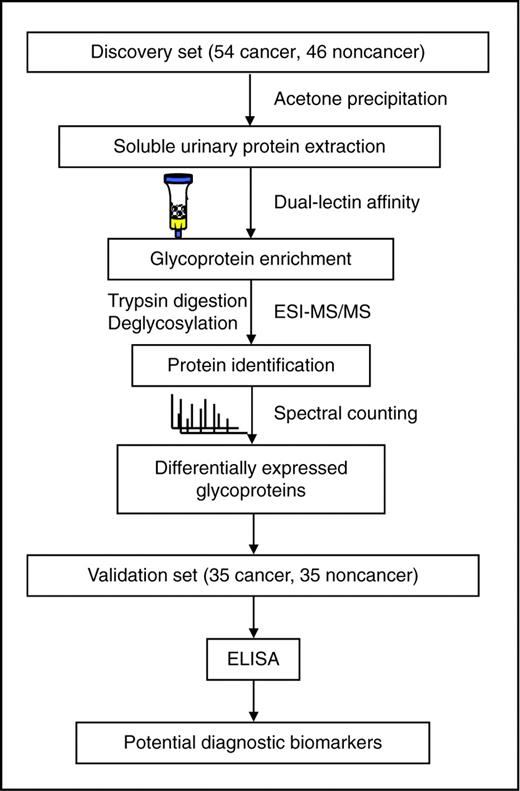 Workflow of LC/MS-MS–based identification and immunoassay-based validation.
Workflow of LC/MS-MS–based identification and immunoassay-based validation.
Results:
Identification of glycoproteins: A total of 267 distinct glycoproteins were identified, with 100 selected for further analysis based on variability.
Technical reproducibility: High reproducibility was observed in LC/MS-MS analysis, with correlation coefficients indicating minimal technical variation.
Differential expression analysis: Three glycoproteins were found to be significantly upregulated in bladder cancer samples, including A1AT.
Validation by Western blotting and ELISA: A1AT was confirmed to be elevated in bladder cancer samples, with potential diagnostic utility demonstrated through ROC curve analysis.
Future implications: The study highlights the potential of urinary glycoprotein profiling as a noninvasive diagnostic tool for bladder cancer and underscores the need for larger confirmatory studies.
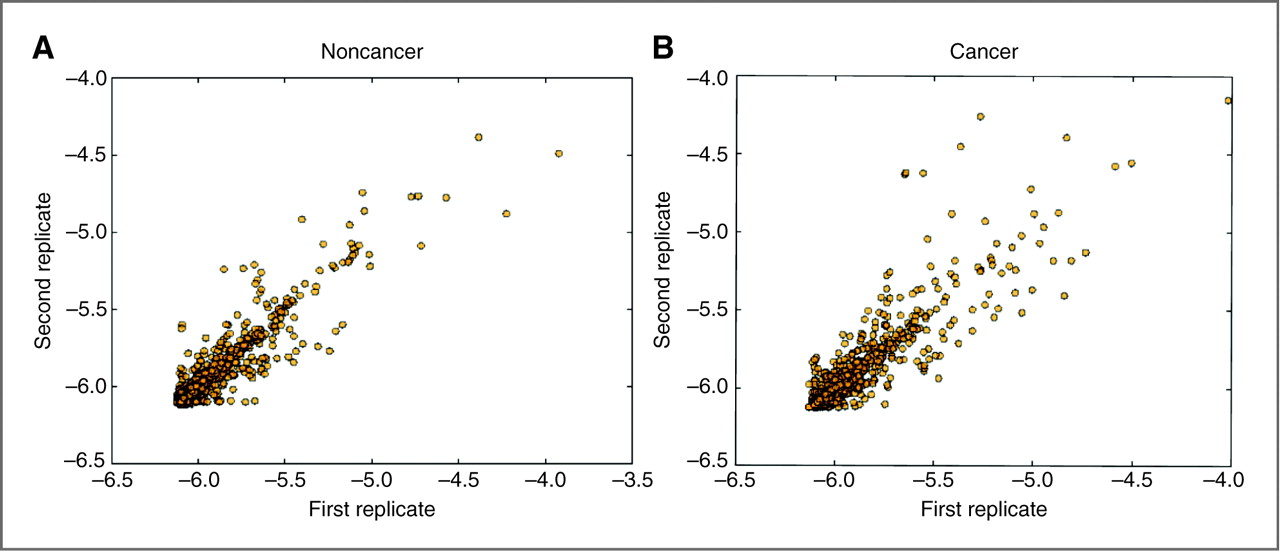 Scatter plots of LTQ replicates across all urine samples based on transformed spectral counts.
Scatter plots of LTQ replicates across all urine samples based on transformed spectral counts.
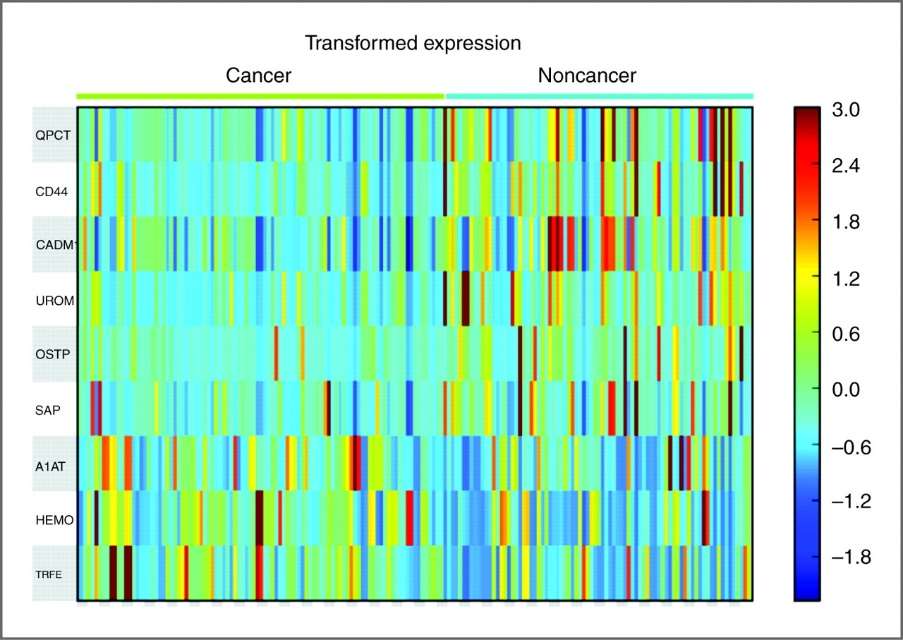 Heat map of differentially expressed urinary glycoproteins identified in duplicate analyses of 103 samples.
Heat map of differentially expressed urinary glycoproteins identified in duplicate analyses of 103 samples.
Reference
- Yang, Na, et al. "Urinary glycoprotein biomarker discovery for bladder cancer detection using LC/MS-MS and label-free quantification." Clinical Cancer Research 17.10 (2011): 3349-3359.




