- Service Details
- Case Study
What is Glycan Methylation Modification?
Glycan methylation is a biochemical process that involves the addition of methyl groups to glycans. Glycans are complex carbohydrates composed of sugar molecules linked together in specific arrangements. Methylation, in the context of glycans, refers to the attachment of a methyl group (CH3) to particular sites on these carbohydrate structures.
Glycans are crucial components in various biological processes, serving roles in cell adhesion, signaling, and molecular recognition. The addition of methyl groups to glycans is a post-translational modification, meaning it occurs after the glycan is initially synthesized and attached to a protein or lipid.
The process of glycan methylation is highly dynamic and regulated. It can occur on specific sugar residues within the glycan structure. This modification influences the overall structure and properties of glycans, impacting their interactions with other molecules and their biological functions.
Understanding glycan methylation is essential in the field of glycobiology, as it provides insights into the intricate mechanisms governing cellular processes. Researchers study glycan methylation to unravel its role in health and disease, as alterations in glycan structures can have significant implications for various physiological and pathological conditions.
Technical Platform for Glycan Methylation Analysis
- Sample Preparation: Obtain glycan-containing samples from glycoproteins or glycolipids. Extract and purify glycans using enzymatic digestion or chemical extraction methods.
- Derivatization: Perform derivatization to enhance detection sensitivity and chromatographic properties. Methylation involves adding methyl groups to glycan hydroxyl groups.
- Chromatographic Separation: Utilize chromatographic techniques like high-performance liquid chromatography (HPLC) or ultra-high-performance liquid chromatography (UHPLC) for separation based on glycan properties.
- Mass Spectrometry (MS) Analysis: Employ mass spectrometry for accurate mass determination and structural elucidation of methylated glycans. Common techniques include matrix-assisted laser desorption/ionization (MALDI) or electrospray ionization (ESI).
- Data Processing and Analysis: Process raw mass spectrometry data to obtain glycan profiles. Analyze methylation patterns, including the number and positions of methyl groups on specific glycan structures.
- Quantitative Analysis: Conduct quantitative analysis to determine the relative abundance of different methylated glycan species. Use standards or internal standards for accurate quantification.
- Bioinformatics and Interpretation: Employ bioinformatics tools to interpret the glycan modification data. Relate methylation patterns to biological functions or disease states.
- Quality Control: Implement quality control measures throughout the analysis to ensure reproducibility and reliability of results. Include appropriate standards and controls in each analytical run.
- Reporting and Documentation: Compile and report findings, including glycan structures, methylation patterns, and quantitative data. Document experimental procedures, parameters, and conditions for reproducibility.
Sample Requirements for Glycan Methylation Analysis
| Sample Type | Sample Size Recommendation |
|---|---|
| Glycoprotein | 50-500 micrograms |
| Glycolipid | 10-100 micrograms |
| Serum/Plasma | 10-100 micrograms |
| Tissues | 5-50 milligrams |
| Cultured Cells | 10-100 micrograms |
| Saliva | 50-500 micrograms |
| Urine | 50-500 micrograms |
| Cerebrospinal Fluid | 50-500 micrograms |
| Synovial Fluid | 50-500 micrograms |
| Cell Culture Media | 1-10 milliliters |
Case. Functional Characterization of Lb-Tec2 and Its Interaction with C. elegans Reveals Insights into Glycan-Mediated Biotoxicity
Background:
This study aims to investigate the functional properties of Lb-Tec2, a protein from Laccaria bicolor, and its impact on C. elegans. The focus is on understanding the glycan-mediated biotoxicity mechanism, involving bacterial strains, nematodes, and recombinant proteins.
Samples:
- Bacterial Strains: E. coli DH5α, BL21(DE3), ArcticExpress (DE3), and specific O8, O9a strains.
- C. elegans Strains: Wild-type (N2) and various mutants with distinct genotypes.
Technical Method:
Cloning and Expression: Lb-Tec2 ORF amplified from L. bicolor genomic DNA. Limulus lectin L6 cDNA ordered and cloned into pET24a vector. Expression in E. coli BL21(DE3) and ArcticExpress (DE3).
Purification of Recombinant Lb-Tec2: Cell collection, lysis, Sepharose column binding, elution with GlcNAc. Stability tested in Hepes and Tris buffers.
C. elegans Biotoxicity Assays: Liquid toxicity assays using E. coli expressing Lb-Tec2 or limulus lectin L6. Fraction calculation from five independent experiments.
TAMRA Labeling and Fluorescence Microscopy: Lb-Tec2 labeled with TAMRA succinimidyl ester. Fluorescence microscopy of C. elegans using labeled protein.
Bacterial Agglutination Assay: E. coli strains agglutination assays with Lb-Tec2. Negative control using BSA.
Structural Analysis of C. elegans N- and O-Glycans: Mass spectrometry for detailed structural analysis.
Monosaccharide Analysis of C. elegans N-Glycans: HPLC used for monosaccharide analysis.
Microcalorimetry: Isothermal titration calorimetry performed on MicroCal iTC200.
Phylogenetic Analysis: Amino acid sequences analyzed for constructing a phylogenetic tree.
Chemical Synthesis: Synthesis of allyl-mannosides and -fucosides for the study.
Statistical Analysis: Mann–Whitney U test used for statistical significance in toxicity assays and monosaccharide analysis.
Results:
- Lb-Tec2, expressed and purified successfully, showed stability in different buffers.
- C. elegans biotoxicity assays revealed insights into glycan-mediated toxicity.
- TAMRA-labeled Lb-Tec2 enabled visualization in C. elegans.
- Bacterial agglutination assays demonstrated interactions with specific E. coli strains.
- Detailed structural and monosaccharide analyses provided insights into glycan composition.
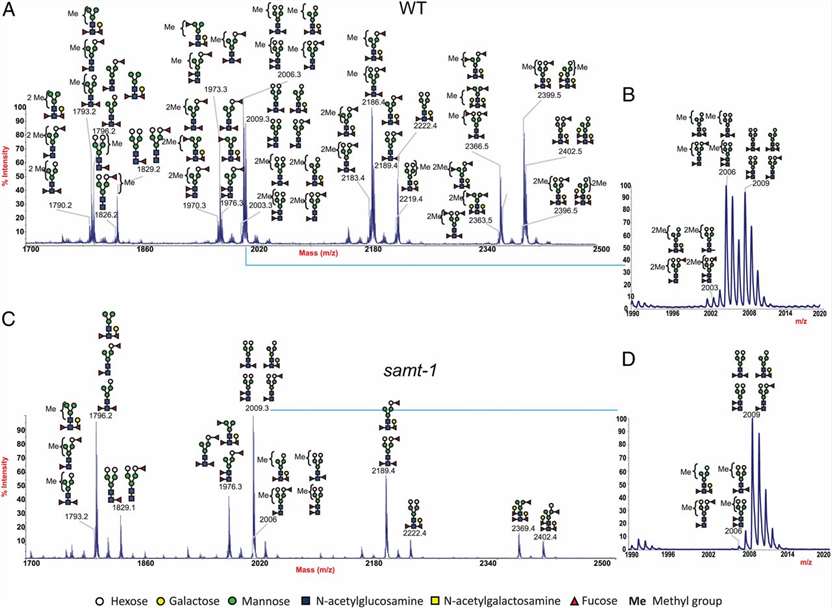 MALDI-TOF spectra of deuteromethylated wild-type and mutant C. elegans N-glycans released by PNGase A digestion.
MALDI-TOF spectra of deuteromethylated wild-type and mutant C. elegans N-glycans released by PNGase A digestion.
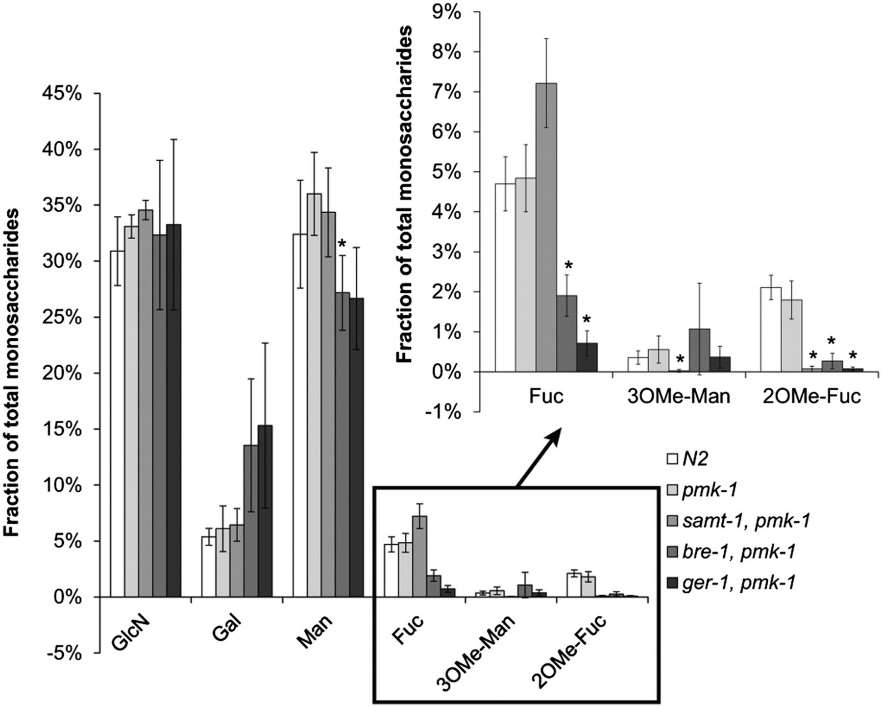 Monosaccharide composition of N-glycans from Lb-Tec2–sensitive and –resistant C. elegans strains.
Monosaccharide composition of N-glycans from Lb-Tec2–sensitive and –resistant C. elegans strains.
Reference
- Wohlschlager, Therese, et al. "Methylated glycans as conserved targets of animal and fungal innate defense." Proceedings of the National Academy of Sciences 111.27 (2014): E2787-E2796.




