- Service Details
- Case Study
What is Lipopolysaccharide?
Lipopolysaccharide (LPS), also known as endotoxin, is a complex molecule found mainly in the outer membrane of Gram-negative bacteria. Structurally, LPS consists of three main components: lipid A, core oligosaccharide and O antigenic polysaccharide chain. These molecules play key roles in bacterial physiology, virulence and immune evasion mechanisms.
Understanding the structural composition of LPS can help to elucidate its biological activity and immunological properties. Analytical techniques such as mass spectrometry and chromatography allow for the precise identification and quantification of LPS components, providing insight into bacterial classification, virulence factors and antibiotic resistance mechanisms.
Lipopolysaccharides have potent immunostimulatory properties and are capable of triggering strong inflammatory responses in mammalian hosts. Analysis of lipopolysaccharide-induced immune responses and endotoxins will help to understand host-pathogen interactions and develop targeted therapeutic agents and vaccines.
In addition, elucidating the structure of the LPS molecule can aid in vaccine design and development. By deciphering the three-dimensional structure of LPS and its interaction with host receptors, researchers can design effective vaccines against bacterial pathogens and reduce the risk of infectious diseases.
Creative Proteomics offers a variety of lipopolysaccharide analysis services to meet your different research needs. Please feel free to contact us if you have any questions.
Lipopolysaccharide Analysis Services by Creative Proteomics
LPS Composition Analysis
Unraveling the complex composition of lipopolysaccharides is critical to understanding their biological activity and immunomodulatory effects, and Creative Proteomics offers comprehensive LPS compositional analysis using advanced analytical techniques, including lipid A structure determination, core oligosaccharide analysis, and O antigen characterization. Our high-resolution mass spectrometry platform, combined with advanced chromatographic methods, enables precise identification and quantification of LPS components, providing insight into bacterial physiology and pathogenesis.
Immunological analysis of LPS
The immunomodulatory properties of lipopolysaccharide highlight its importance in host-pathogen interactions and inflammatory responses. We offer a suite of immunological assays for elucidating the immunogenicity, endotoxicity, and immune activation characteristics of LPS molecules. From cytokine analysis and cellular assays to endotoxin quantification, our comprehensive immunoassays enable researchers to dissect the intricate immune cascade triggered by LPS exposure, thereby facilitating the development of novel therapies and vaccines.
Structural elucidation of LPS
The structural diversity of lipopolysaccharides poses challenges in deciphering their functional roles and antigenic properties. We combined spectroscopic techniques (e.g., nuclear magnetic resonance (NMR) spectroscopy and X-ray crystallography) and computational modeling to reveal the three-dimensional structure of the LPS molecule. By elucidating the spatial arrangement of lipid A, core oligosaccharide, and O antigenic molecules, our structural analysis provides insights into LPS-host receptor interactions, bacterial pathogenesis, and vaccine design strategies.
Technical Platforms for Lipopolysaccharide Analysis
Mass Spectrometry (MS): High-resolution mass spectrometry enables accurate identification and quantification of LPS constituents, facilitating compositional analysis and structural elucidation.
Chromatographic Techniques: Liquid chromatography (LC) and gas chromatography (GC) coupled with mass spectrometry allow for separation and characterization of LPS components based on their physicochemical properties.
Immunological Assays: Enzyme-linked immunosorbent assays (ELISA), flow cytometry, and cytokine profiling techniques enable assessment of LPS-induced immune responses and endotoxicity.
Spectroscopic Methods: Nuclear magnetic resonance (NMR) spectroscopy, infrared (IR) spectroscopy, and circular dichroism (CD) spectroscopy provide insights into the structural conformation and dynamics of LPS molecules.
Computational Modeling: Molecular modeling and bioinformatics tools aid in predicting LPS structure-function relationships and simulating LPS-receptor interactions.
Sample Requirements for Lipopolysaccharide Analysis
| Sample Type | Sample Volume | Sample Preparation |
|---|---|---|
| Bacterial Culture | 1 mL | Centrifuge at 10,000 × g for 10 minutes to pellet cells. |
| Resuspend pellet in 1 mL phosphate-buffered saline (PBS). | ||
| Heat at 100°C for 10 minutes to lyse cells. | ||
| Centrifuge at 10,000 × g for 10 minutes to remove debris. | ||
| Serum/Plasma | 100 µL | Dilute serum/plasma 1:10 in PBS. |
| Heat at 70°C for 10 minutes to inactivate complement. | ||
| Centrifuge at 10,000 × g for 10 minutes to remove debris. | ||
| Tissue Homogenate | Variable | Homogenize tissue in 1 mL PBS per 100 mg tissue. |
| Centrifuge at 10,000 × g for 10 minutes to remove debris. | ||
| Collect supernatant for analysis. | ||
| Environmental Sample | Variable | Filter environmental sample through 0.22 µm filter. |
| Dilute filtrate as needed in PBS. | ||
| Centrifuge at 10,000 × g for 10 minutes to remove debris. | ||
| Collect supernatant for analysis. |
Applications of Lipopolysaccharide Analysis
Microbiology and Infectious Diseases: LPS analysis is crucial for identifying and characterizing bacteria, especially Gram-negative bacteria, which possess LPS in their outer membrane. Different bacterial strains can be differentiated based on variations in their LPS structure, aiding in the diagnosis and treatment of infectious diseases.
Vaccine Development: Understanding the structure and composition of LPS is essential for developing vaccines against bacterial infections. LPS molecules are potent immunostimulants and are often used as adjuvants in vaccine formulations to enhance the immune response.
Endotoxin Detection: LPS is also referred to as endotoxin when it is released from bacterial cells and triggers immune responses in mammals. In biomedical research and pharmaceutical manufacturing, it is critical to detect and quantify endotoxin levels in substances such as drugs, medical devices, and biological products to ensure their safety and quality.
Biotechnology and Biomedical Research: LPS analysis is employed in various research areas, including immunology, inflammation, and host-pathogen interactions. Researchers use techniques such as mass spectrometry, chromatography, and immunoassays to study the structure, function, and biological effects of LPS molecules.
Environmental Monitoring: LPS analysis can be utilized in environmental microbiology to assess microbial contamination in soil, water, and air samples. Monitoring LPS levels can help evaluate the presence of potentially harmful bacteria and assess environmental health and safety.
Food Safety and Quality Control: Certain foodborne pathogens, such as Salmonella and Escherichia coli, produce LPS, which can contaminate food products and cause food poisoning. Analyzing LPS levels in food samples can aid in identifying contamination sources, implementing preventive measures, and ensuring food safety.
Drug Development and Therapeutics: LPS-induced inflammation plays a role in various diseases, including sepsis, inflammatory bowel disease, and autoimmune disorders. Understanding the mechanisms of LPS-mediated inflammation can facilitate the development of novel therapeutic strategies and drugs targeting inflammatory pathways.
Title: Lipid A Variants in P. multocida Isolate PM2 and Their Implications in Haemorrhagic Septicaemia
Background
Haemorrhagic septicaemia (HS) is associated with P. multocida infections in cloven-hoofed ungulates, presenting in acute, sub-acute, and chronic forms. Despite the strong link between HS and the endotoxicity of lipid A, the structural elucidation of lipid A in P. multocida remains unreported.
Sample
The study focused on P. multocida isolate PM2 obtained from Veterinary Research Institute, Peshawar, and subjected to bacterial identification and LPS extraction.
Technical Methods
Bacterial Identification: PM2 isolate was confirmed using PCR targeting specific genes (KMT1, 6b, bcbD) associated with P. multocida and its serotype.
LPS Extraction and Purification: PM2 cells were grown, treated with formalin, and harvested for LPS extraction using the hot-phenol method. Extracted LPS were purified, dialyzed, and analyzed by DOC-PAGE.
Acid Hydrolysis and Extraction of Lipid A: Purified LPS underwent mild acid hydrolysis to extract lipid A. The hydrolyzed solution was centrifuged, and the pellet was dissolved in chloroform:methanol:water. The chloroform extract was dried, redissolved, and used for ESI-MS analysis.
Mass Spectrometry of Lipid A: ESI-MS/MS was conducted using LTQ XL mass spectrometer in negative ionization mode. Full scan mass spectrum data was acquired, and ions were fragmented by CID. Data analysis was performed using Xcalibur and Chemdraw software.
Results
Complexity and Heterogeneity of Lipid A: ESI-MS analysis revealed a diverse range of lipid A variants in PM2, ranging from hepta-acylated to tetra-acylated, with varied decorations of Ara4N moiety on the glucosamine backbone.
Structural Characteristics: Hepta-acylated, hexa-acylated, penta-acylated, and tetra-acylated lipid A variants exhibited specific acylation patterns and Ara4N placement, influencing their functional properties.
Functional Implications: Structural variations in lipid A, including palmitoylation and Ara4N placement, contribute to evasion of host immune responses and potentially influence the pathogenicity of P. multocida.
Role of Enzymes: Enzymes such as PagP and PagL are implicated in acyl chain addition and removal, respectively, influencing lipid A structure and host interactions.
Immunostimulatory Activities: Bisphosphorylated hexa-acylated and hepta-acylated lipid A variants induce high inflammatory responses, potentially playing a pivotal role in HS pathogenesis.
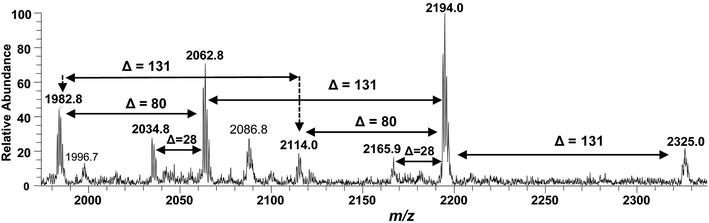 ESI-MS full scan of lipid A from PM2, in negative ionization mode, showing the correlation among hepta-acylated lipid A variants.
ESI-MS full scan of lipid A from PM2, in negative ionization mode, showing the correlation among hepta-acylated lipid A variants.
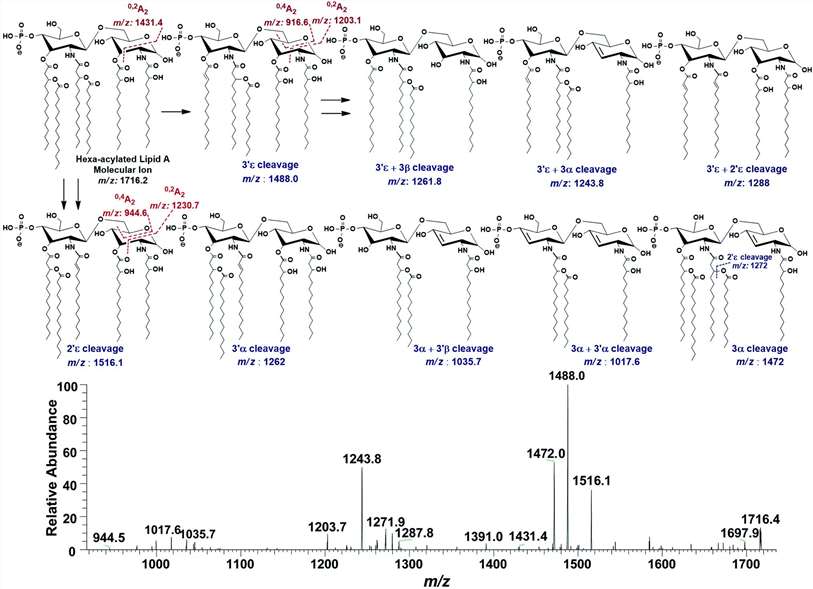 (A) ESI-MS2 of hexa-acylated lipid A molecular ion at m/z 1756.3 @CID 47, showing its fragmentation patterns, (B) MS3 of base peak at m/z 1528.5 @CID 22, obtained from MS2 of m/z 1756.3.
(A) ESI-MS2 of hexa-acylated lipid A molecular ion at m/z 1756.3 @CID 47, showing its fragmentation patterns, (B) MS3 of base peak at m/z 1528.5 @CID 22, obtained from MS2 of m/z 1756.3.
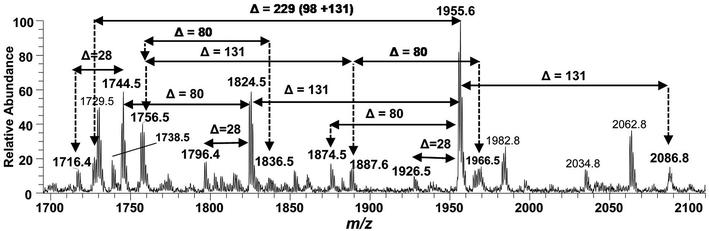 Negative ion mode ESI-MS full scan of lipid A from PM2, showing the correlation among hexa-acylated lipid A variants.
Negative ion mode ESI-MS full scan of lipid A from PM2, showing the correlation among hexa-acylated lipid A variants.
Reference
- Tawab, Abdul, et al. "Mass spectrometric analysis of lipid A obtained from the lipopolysaccharide of Pasteurella multocida." RSC advances 10.51 (2020): 30917-30933.




