- Service Details
- Case Study
What is Glycoprotein Amino Acid Sequence Analysis?
Glycoproteins are essential constituents of living organisms, playing pivotal roles in various biological processes such as cellular recognition, signal transduction, and immune responses. These macromolecules are characterized by the presence of oligosaccharide chains covalently linked to specific amino acid residues, primarily serine, threonine, or asparagine. The intricate interplay between the protein and carbohydrate moieties confers unique structural and functional properties to glycoproteins, influencing their stability, solubility, and interactions with other biomolecules.
The amino acid sequence forms the backbone of glycoproteins, determining their overall structure and functionality. Unlike simple proteins, glycoproteins exhibit significant heterogeneity in their amino acid sequences due to variations in glycosylation patterns and sites. Glycoprotein amino acid sequence analysis involves investigating the composition and sequence of amino acids within glycoproteins, including the identification of specific amino acids and their properties. The amino acid sequence provides crucial insights into the structure, function, and interactions of proteins, serving as a fundamental basis for understanding their biological roles.
Specific Glycoprotein Amino Acid Sequence Analysis Projects Offered by Creative Proteomics
At Creative Proteomics, we offer services tailored to your specific research needs. We undertake a broad spectrum of Glycoprotein Amino Acid Sequence Analysis tasks:
- Sequence Analysis: We offer full sequence coverage to confirm the identity and evaluate the integrity of your glycoprotein samples.
- Site Occupancy Determination: Our team quantifies the extent of glycosylation on a specific protein site.
- Glycan Structure Analysis: We analyze the complex carbohydrate structures of your glycoproteins. This includes profiling and site-specific analysis.
- Qualitative and Quantitative Glycomics and Glycoproteomics: We perform extensive studies characterizing glycome and glycoproteome changes across different biological states.
- Data Analysis: We employ sophisticated bioinformatics tools to interpret the mass spectrometric data, allowing you to derive meaningful insights from your analysis.
- Validation: We conduct a thorough validation of the analysis results, ensuring the highest level of accuracy and reliability.
Techniques and Platforms for Glycoprotein Amino Acid Sequence Analysis
At Creative Proteomics, we are equipped with a state-of-art technical platform that is specifically designed and established to perform comprehensive glycoprotein amino acid sequence analysis. Technically, a combination of high-resolution mass spectrometry (MS) and liquid chromatography (LC) forms the backbone of our analytical platform.
High-resolution MS, including MALDI-TOF MS and ESI-MS, is employed for high-sensitivity detection and accurate mass determination of glycopeptides. These techniques are complemented by LC, which allows for efficient separation and simplification of complex glycopeptide mixtures ahead of MS analysis.
We have also developed advanced bioinformatics tools and software, enabling accurate identification of peptides and glycans, annotation of glycosylation sites, and prediction of glycan structures based on MS data.
Importantly, our platform also integrates multiple orthogonal techniques for validation of the MS-based results. For instance, Edman sequencing and western blotting can be added to validate the identified glycosylation sites and the expression levels of specific glycoproteins, respectively.
Sample Requirements for Glycoprotein Amino Acid Sequence Analysis
| Sample Type | Sample Amount |
|---|---|
| Serum | 100-200 μL |
| Plasma | 100-200 μL |
| Cell Culture Medium | 1-5 mL |
| Tissue Homogenate | 50-100 mg |
| Purified Protein | 10-50 μg |
| Cell Lysate | 1-5 x 106 cells |
| Biopsy | 2-5 mm3 |
| Saliva | 100-500 μL |
| Urine | 1-5 mL |
| CSF (Cerebrospinal Fluid) | 100-200 μL |
| Synovial Fluid | 100-200 μL |
| Amniotic Fluid | 100-200 μL |
| Breast Milk | 100-500 μL |
| Feces | 50-100 mg |
Case. Glycoproteomic Profiling Reveals Potential Serum Biomarkers for Narcolepsy Type 1
Background:
Narcolepsy Type 1 (NT1) is a neurological disorder characterized by excessive daytime sleepiness and disrupted sleep patterns. The exact etiology of NT1 remains incompletely understood, highlighting the need for further research into its underlying molecular mechanisms. Aberrant glycosylation patterns in serum proteins have been implicated in various diseases, suggesting that glycoproteomic profiling could unveil potential biomarkers for NT1.
Samples:
The study utilized serum samples from both NT1 patients and healthy controls. A total of 27 serum samples were analyzed, including 16 NT1 samples and 11 healthy controls. Samples were subjected to depletion of high-abundance proteins using a Human 14 Multiple Affinity Removal Column to enrich low-abundance glycoproteins for glycoproteomic analysis.
Technical Method:
Depletion and Enrichment: High-abundance proteins were removed using a Human 14 Multiple Affinity Removal Column to expand the dynamic range of low-abundance serum proteins. Enrichment of glycopeptides was achieved through hydrophilic interaction liquid chromatography (HILIC), enhancing the identification of N-glycopeptides.
LC-MS/MS Analysis: Depleted and enriched samples underwent liquid chromatography-tandem mass spectrometry (LC-MS/MS) analysis. Samples were divided into two parts: one analyzed without enrichment and the other subjected to HILIC enrichment prior to LC-MS analysis. Data-dependent acquisition (DDA) mode was employed to identify N-glycopeptides present in the samples.
Data Analysis: Glyco-Decipher software was utilized for preliminary analysis of the samples' raw files. Structures of glycopeptides were manually confirmed based on various parameters, including monoisotopic mass, charge, retention time, and MS2 spectra. Statistical analysis, including Mann–Whitney U-test and receiver operating characteristic (ROC) curve analysis, was performed to identify significant glycopeptides differentiating between NT1 and control samples.
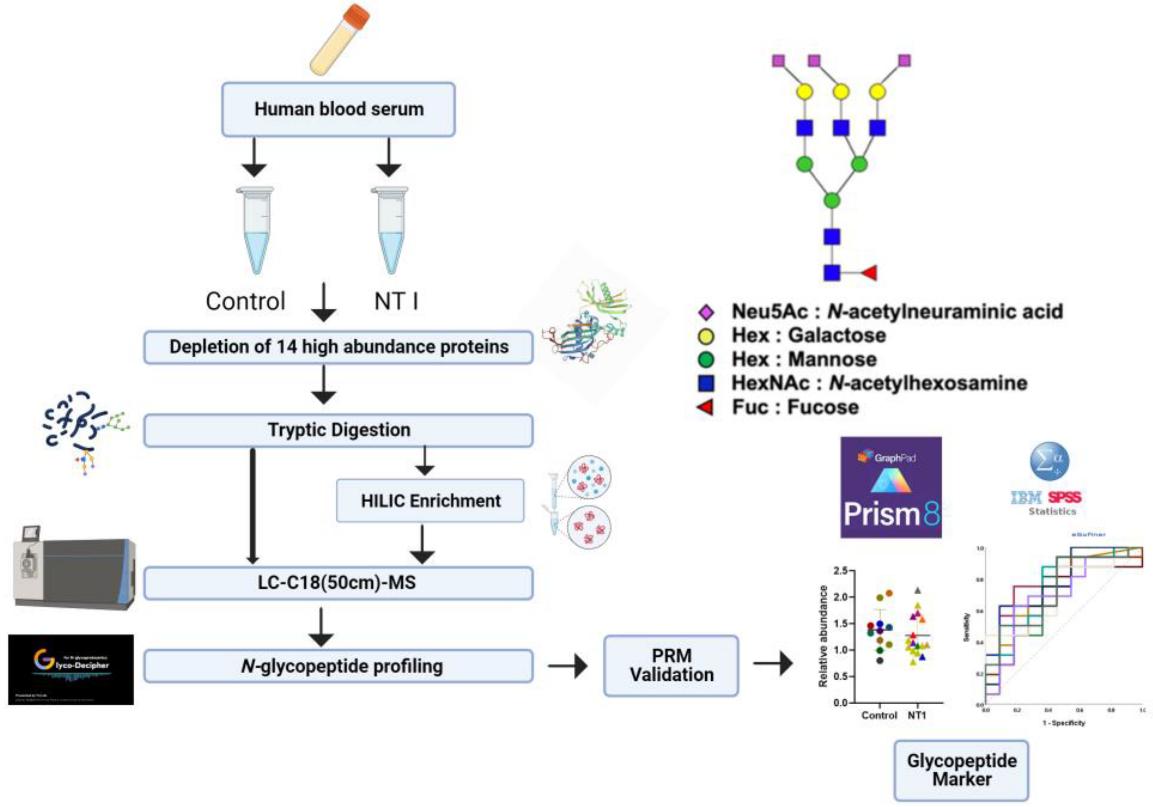 Workflow for LC-MS/MS analysis.
Workflow for LC-MS/MS analysis.
Results:
Identification of Significant Glycopeptides: The study identified 28 significant glycopeptides differentially expressed between NT1 and control samples. These glycopeptides, derived from proteins such as inter-alpha-trypsin inhibitor H4, complement factor B, ceruloplasmin, fibronectin, and prothrombin, have potential as biomarkers for NT1.
Glycan Heterogeneity and Abundance: Sialylated glycopeptides were found to be the most abundant, with a decrease in their abundance observed in NT1 samples. HILIC enrichment significantly enhanced the identification of glycopeptides, showing glycan heterogeneity in the samples.
Validation of Biomarkers: A targeted parallel reaction monitoring (PRM) approach validated significant glycopeptides, confirming their fold change trends observed in untargeted analysis. Seven glycopeptides showed consistent fold change trends between NT1 and control samples.
Clinical Implications: The study suggests that assessing serum N-glycopeptides could aid in early diagnosis and risk assessment of NT1, potentially guiding the development of novel therapeutic strategies targeting aberrant glycosylation patterns.
Biological Insights: The identified glycopeptides shed light on potential molecular mechanisms underlying NT1, emphasizing the role of glycosylation in immunological activities and inflammation associated with the disorder.
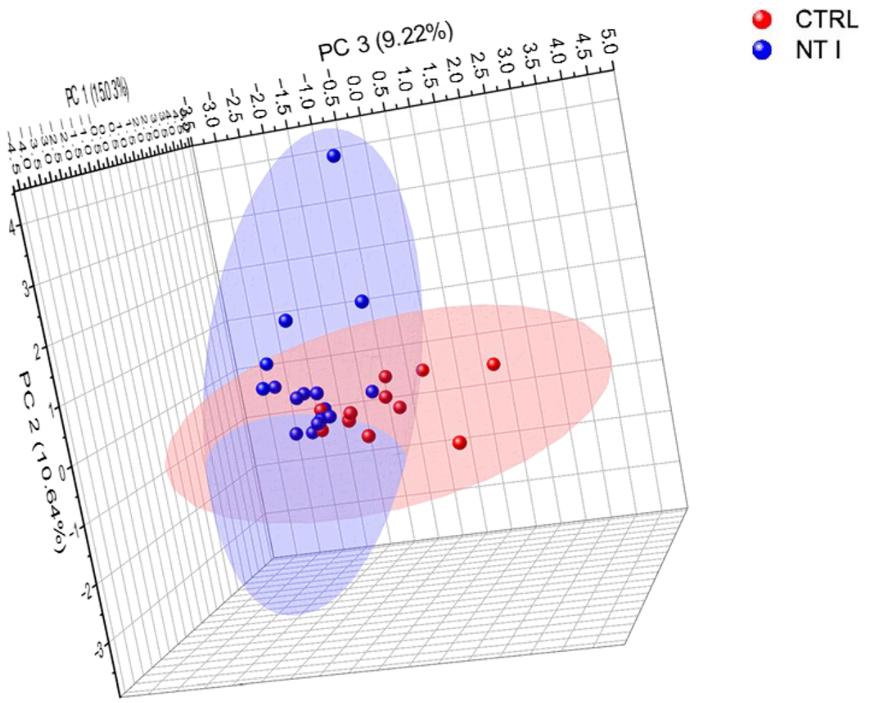 Unsupervised principal component analysis (PCA) at 95% confidence level for total identified glycopeptides.
Unsupervised principal component analysis (PCA) at 95% confidence level for total identified glycopeptides.
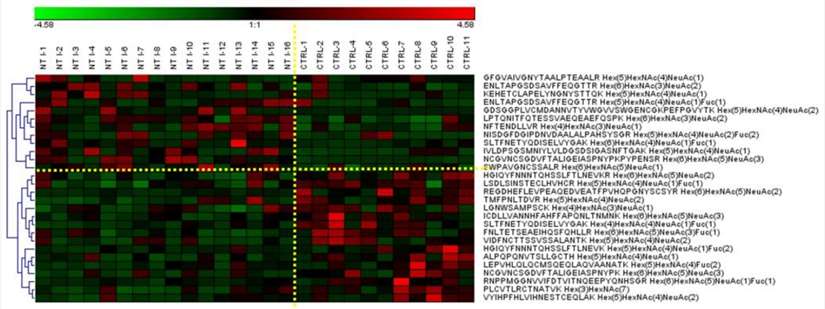 Heatmap of N-glycopeptides with statistically significant expression structures between control and NT1 samples.
Heatmap of N-glycopeptides with statistically significant expression structures between control and NT1 samples.
Reference
- Ivleva, Vera B., et al. "Overcoming challenges in structural characterization of HIV-1 envelope glycoprotein by LC-MS/MS." Journal of The American Society for Mass Spectrometry 30.9 (2019): 1663-1678.




