- Service Details
- Case Study
What is Stable Isotope Quantification Analysis?
Stable Isotope Quantification is a powerful and precise technique in the field of mass spectrometry. This method allows for accurate quantification of targeted molecules by labeling them with non-radioactive, stable isotopes. In the context of glycoprotein analysis, stable isotopes are used to label glycopeptides, enabling not only the identification of the glycosites but their quantification as well. This application provides deeper insights into the variables influencing protein function, including disease states and therapeutic responses.
Technology Platform for Glycoprotein Stable Isotope Quantification Analysis
With the advent of cutting-edge proteomic technologies, the analysis and quantification of glycoproteins have become significantly easier. Creative Proteomics offers one such advanced platform for stable isotope quantification of glycoproteins.
Isotope-Coded Affinity Tags (ICAT)
One of the most critical technologies employed at Creative Proteomics for stable isotope quantification of glycoproteins is the Isotope-Coded Affinity Tags (ICAT) approach. It is based on the principle of labeling peptides with isotopes, which differ in their mass, but not in their chemical properties. The advantage of this approach is that it allows quantitative analysis of glycoproteins as each protein's isotope abundance can be measured accurately using mass spectrometry.
Liquid Chromatography-Mass Spectrometry (LC-MS)
Liquid Chromatography-Mass Spectrometry (LC-MS) is also used for stable isotope quantification of glycoproteins. LC-MS combines the physical separation capabilities of liquid chromatography with the mass analysis capabilities of mass spectrometry. It can provide a comprehensive view of the glycoprotein signature within a sample as it allows for high-resolution separation of complex samples and high sensitivity and accuracy in protein identification and quantification.
Tandem Mass Tag (TMT) Labeling
At Creative Proteomics, Tandem Mass Tag (TMT) labeling is also adopted for glycoprotein stable isotope quantification. This method involves labeling the N-terminus and side-chain amines of peptides with tags that can be cleaved in the mass spectrometer, allowing for simultaneous quantification of multiple samples in a single experiment. It leads to enhanced throughput and minimized sample variability, thus providing more reliable quantification results.
18O/16O Labeling
It involves the use of water enriched with the stable heavy oxygen isotope (18O) to exchange the terminal glycans' oxygen in glycoproteins, facilitating relative and absolute quantification.
Dimethyl Labeling
It enables the precise quantification of changes in the amounts of specific glycoproteins in two different samples.
Data Analysis with Latest Bioinformatic Tools
In addition to the advanced experimental techniques, Creative Proteomics also has a powerful bioinformatic infrastructure to handle the vast amount of data generated from glycoprotein stable isotope quantification analyses. Our data scientists utilize the latest bioinformatic tools to match the mass spectrometry data with protein databases accurately, thereby identifying and quantifying the detected proteins.
By utilizing the technological platform, we can harness the power of stable isotope quantification to gain a fuller understanding of the complex mechanisms underlying various biological processes and diseases.
Sample Requirements for Glycoprotein Stable Isotope Quantification Analysis
| Sample Type | Sample Volume | Additional Requirements |
|---|---|---|
| Serum/Plasma | 100-500 μL | EDTA or heparin anticoagulants, centrifugation |
| Cell Culture Supernatant | 1-5 mL | Cell debris removal, protein concentration |
| Tissue Homogenate | 50-200 mg tissue | Homogenization, protein extraction |
| Urine | 1-10 mL | Centrifugation, removal of particulates |
| Cerebrospinal Fluid | 100-500 μL | Centrifugation, removal of cells and debris |
| Saliva | 500 μL - 1 mL | Centrifugation, removal of mucins and debris |
| Biopsy Specimens | Variable | Tissue homogenization, protein extraction |
| Cell Lysates | Variable | Protein extraction, removal of cellular debris |
| Synovial Fluid | 100-500 μL | Centrifugation, removal of cells and debris |
Applications of Glycoprotein Stable Isotope Quantification Analysis
- Protein Turnover Studies: Tracking protein degradation and synthesis rates in biological systems.
- Biomarker Discovery: Identifying disease biomarkers by comparing glycoprotein abundance in healthy vs. diseased samples.
- Drug Development: Studying drug pharmacokinetics and biodistribution for therapeutic glycoproteins.
- Glycoprotein Structure-Function Relationships: Investigating the impact of glycosylation on protein structure and function.
- Vaccine Development: Characterizing viral or bacterial glycoproteins for vaccine efficacy.
- Quality Control in Biopharmaceutical Production: Monitoring glycoprotein expression and post-translational modifications for product consistency.
- Metabolic Flux Analysis: Analyzing glycoprotein biosynthesis pathways and metabolic reprogramming.
- Cellular Signaling Studies: Investigating glycoprotein involvement in cellular signaling pathways and signal transduction mechanisms.
- Neuroscience Research: Studying glycoprotein modifications in neuronal cells and their implications in neurodegenerative diseases.
- Plant Biology: Understanding glycoprotein-mediated plant-pathogen interactions and plant defense mechanisms.
- Environmental Monitoring: Detecting glycoprotein biomarkers as indicators of environmental pollutants or contaminants.
- Food and Agriculture: Assessing glycoprotein profiles in food products for quality control and nutritional analysis.
Case. Glycopeptide Analysis for Profiling Cell Surface Proteins
Background:
Quantitative proteomics plays a pivotal role in functional genomics, aiding in the detection of disease markers, therapeutic targets, and insights into biological processes. Two primary methods, two-dimensional gel electrophoresis (2DE) coupled with mass spectrometry (MS) and stable isotope tagging of proteins with automated peptide MS/MS, have been extensively used for generating quantitative protein profiles. However, both methods face challenges in analyzing the complete proteome due to preferential analysis of abundant proteins. To address this, strategies such as selective isolation of protein subsets and comprehensive analysis of specific sub-proteomes have been employed.
Samples:
Human serum and a crude membrane fraction from the LNCaP prostate cancer epithelial cell line were utilized in the study.
Technical Method:
Glycoprotein oxidation: Carbohydrates in serum glycoproteins are oxidized using sodium periodate.
Coupling: Oxidized glycoproteins are covalently bound to hydrazide groups on a solid support.
Proteolysis: Glycoproteins on the solid support are proteolyzed, and nonglycosylated peptides are removed.
Isotope labeling: Immobilized glycopeptides are labeled with isotopically light or heavy forms of succinic anhydride.
Release: Formerly N-linked glycopeptides are released from the solid phase using PNGase F treatment.
Analysis: The released peptides are identified and quantified using microcapillary high-performance liquid chromatography electrospray ionization (µLC-ESI) MS/MS or µLC separation followed by matrix-assisted laser desorption/ionization (MALDI) MS/MS.
Results:
The glycopeptide capture method efficiently and specifically isolated glycoproteins from human serum, as demonstrated by gel electrophoresis and staining techniques.
Compared to conventional methods, the glycopeptide capture method identified a larger number of serum proteins with reduced sample complexity.
The method demonstrated high selectivity for cell surface proteins, with a majority of the identified glycoproteins being transmembrane proteins.
The glycopeptide capture method enabled the identification of antigens and proteins typically underrepresented in analyses performed using 2DE.
While limitations exist in profiling all serum proteins, the method showed promise for quantitative analysis of glycoproteins, with observed ratios closely matching expected values.
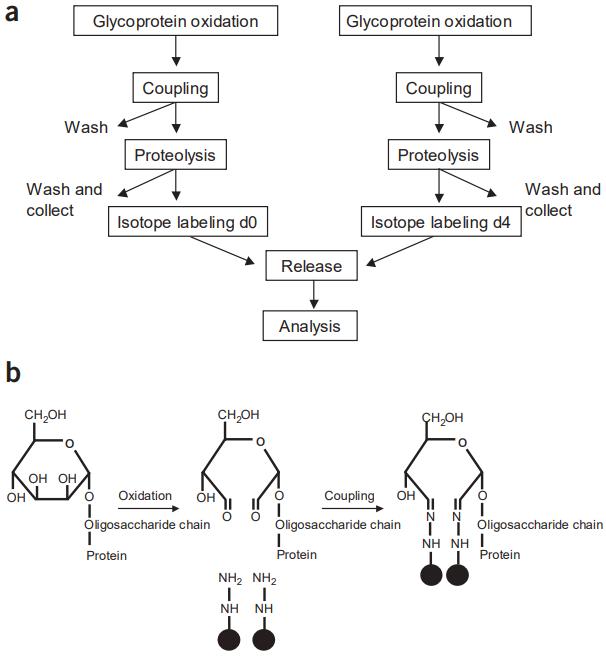 Schematic diagram of quantitative analysis of N-linked glycopeptides.
Schematic diagram of quantitative analysis of N-linked glycopeptides.
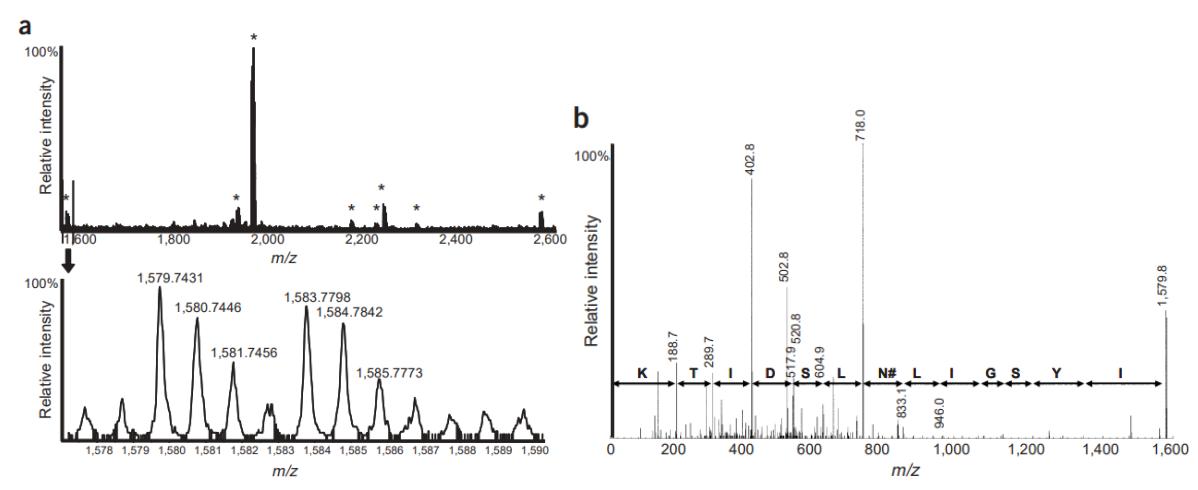 Quantitative analysis of formerly N-linked glycosylated peptides by MALDI QqTOF MS.
Quantitative analysis of formerly N-linked glycosylated peptides by MALDI QqTOF MS.
Reference
- Zhang, Hui, et al. "Identification and quantification of N-linked glycoproteins using hydrazide chemistry, stable isotope labeling and mass spectrometry." Nature biotechnology 21.6 (2003): 660-666.




