What are Sulfatides?
Sulfatides, also known as sulfated galactocerebrosides, are amphipathic molecules primarily found in the myelin sheath of neurons and oligodendrocytes in the central nervous system (CNS), as well as in certain peripheral tissues. Structurally, sulfatides consist of a ceramide molecule, comprising a sphingosine or sphinganine backbone linked to a fatty acid, attached to a sugar moiety, usually galactose or glucose, via a glycosidic bond. Importantly, sulfatides are characterized by the presence of a sulfate group attached to the sugar residue, which imparts a negative charge to the molecule.
The analysis of sulfatides is essential for elucidating their roles in physiological processes and their implications in various diseases. Several neurological disorders, including multiple sclerosis, Alzheimer's disease, and metachromatic leukodystrophy, have been associated with alterations in sulfatide metabolism or levels. Moreover, sulfatides have been implicated in cancer progression and immune modulation, highlighting their diverse functions in health and disease.
Creative Proteomics offers a comprehensive suite of analytical services for sulfatides analysis, leveraging cutting-edge mass spectrometry-based techniques. Our specialized methodologies enable precise identification, quantification, and characterization of sulfatides in various biological samples, including tissues, cells, and biofluids.
Sulfatides Analysis Services by Creative Proteomics
Sulfatides Qualitative Analysis: We employ advanced mass spectrometry techniques to identify sulfatide species present in biological samples with high sensitivity and accuracy.
Sulfatides Quantitative Analysis: Accurate quantification of sulfatides can be achieved through targeted mass spectrometry methods, allowing for the determination of concentration levels in biological samples.
Sulfatides Structural Characterization: Through tandem mass spectrometry and fragmentation analysis, we elucidate the structural details of sulfatide molecules, including fatty acid chain length and saturation, as well as the position of sulfate groups.
Sulfatides Functional Studies: We offer services for investigating the biological functions of sulfatides, including their roles in cellular signaling, lipid metabolism, and disease pathogenesis.
Bioinformatics Analysis: Our bioinformatics team provides comprehensive data analysis and interpretation, facilitating the integration of sulfatide analysis results with relevant biological pathways and networks.
Customized Solutions: We tailor our sulfatide analysis services to meet specific research needs, offering customized experimental designs and data analysis strategies.
Analytical Techniques for Sulfatides Analysis
High-Resolution Mass Spectrometry
High-resolution mass spectrometry (HRMS) serves as the cornerstone of sulfatides analysis, allowing for the accurate determination of molecular masses and structural elucidation. At Creative Proteomics, we harness the power of the following HRMS platforms:
- Thermo Scientific Q Exactive Series: Renowned for its exceptional mass accuracy and resolution, the Q Exactive series enables precise quantification and structural characterization of sulfatide species across various biological samples.
- Waters Xevo TQ-XS: This triple quadrupole mass spectrometer offers exceptional sensitivity and selectivity, making it ideal for targeted quantitative analysis of sulfatides in complex matrices.
Liquid Chromatography (LC) Coupled with Mass Spectrometry
Liquid chromatography coupled with mass spectrometry (LC-MS) enhances the separation and detection of sulfatide species, enabling comprehensive structural characterization and quantitative analysis. At Creative Proteomics, we utilize LC-MS systems in conjunction with HRMS platforms to achieve optimal analytical performance.
Tandem Mass Spectrometry (MS/MS)
Tandem mass spectrometry (MS/MS) facilitates in-depth structural elucidation of sulfatides through fragmentation analysis. By subjecting sulfatide ions to sequential fragmentation events, we unravel the fatty acyl composition, double bond positions, and sulfate group localization, providing valuable insights into sulfatide structure-function relationships.
Matrix-Assisted Laser Desorption/Ionization Imaging Mass Spectrometry (MALDI-IMS)
Matrix-assisted laser desorption/ionization imaging mass spectrometry (MALDI-IMS) enables the visualization of sulfatide distribution within biological tissues with spatial resolution. This powerful technique offers insights into sulfatide localization at the cellular and subcellular levels, facilitating the study of sulfatide biology and pathology in situ.
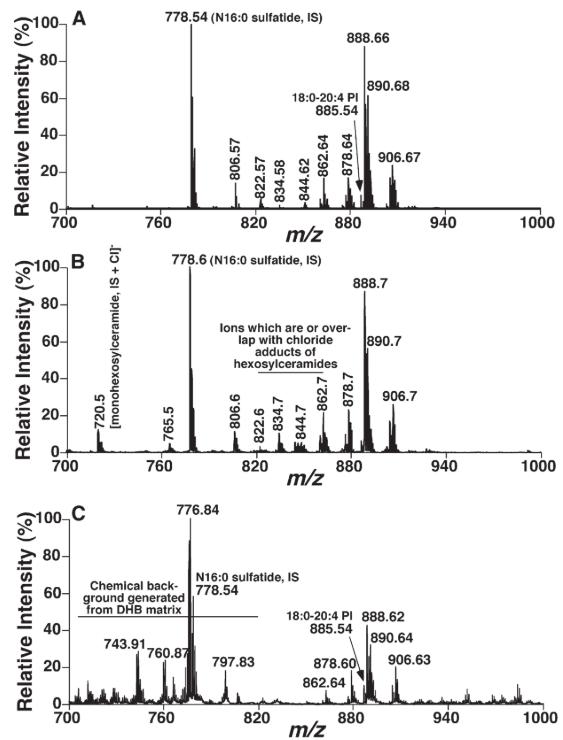 Mass spectral comparison of sulfatide molecular species present in mouse brain cortical lipid extracts acquired by either MALDI-MS or ESI-MS (Cheng et al., 2010).
Mass spectral comparison of sulfatide molecular species present in mouse brain cortical lipid extracts acquired by either MALDI-MS or ESI-MS (Cheng et al., 2010).
Sample Requirements for Sulfatides Analysis
| Sample Type | Recommended Sample Volume |
|---|---|
| Serum/Plasma | 100-200 µL |
| Cerebrospinal Fluid (CSF) | 50-100 µL |
| Tissue (e.g., Brain, Kidney) | 10-50 mg |
| Cell Culture (e.g., Cells, Tissues) | 106 - 107 cells |
| Urine | 1-2 mL |
| Milk | 1-2 mL |
| Saliva | 100-200 µL |
| Synovial Fluid | 50-100 µL |
| Amniotic Fluid | 50-100 µL |
| Feces | 100-200 mg |
Deliverables for Sulfatides Analysis
- Analytical Report: Detailed summary of experimental procedures, methods used, and results obtained.
- Quantitative Data: Sulfatide concentrations with calibration curves and quality control results.
- Structural Characterization: Information on sulfatide species identified, including fatty acyl composition and sulfate group localization.
- Visualization Data: Imaging mass spectrometry results depicting sulfatide distribution in tissues (if applicable).
- Interpretation and Discussion: Insights into the biological significance of results and potential implications.
Applications of Sulfatides Analysis
Neuroscience Research: Sulfatides are abundant in the central nervous system and are implicated in myelin stability, neuronal signaling, and synaptic function. Sulfatides analysis facilitates the study of neurodegenerative disorders such as multiple sclerosis, Alzheimer's disease, and Parkinson's disease, providing insights into disease mechanisms and potential therapeutic targets.
Cancer Biology: Dysregulation of sulfatide metabolism has been observed in various cancers, including breast cancer, ovarian cancer, and prostate cancer. Sulfatides analysis enables the characterization of sulfatide profiles in cancer cells and tissues, contributing to the understanding of tumor progression, metastasis, and drug resistance mechanisms.
Metabolic Disorders: Inherited metabolic disorders such as metachromatic leukodystrophy and Krabbe disease are characterized by deficiencies in sulfatide metabolism enzymes. Sulfatides analysis aids in the diagnosis and monitoring of these disorders by assessing sulfatide levels in biological samples such as blood, urine, and cerebrospinal fluid.
Cardiovascular Disease: Sulfatides have been implicated in cardiovascular health, with studies suggesting their involvement in atherosclerosis and thrombosis. Sulfatides analysis allows for the investigation of sulfatide-mediated signaling pathways and lipid-lipid interactions in vascular endothelial cells and platelets, shedding light on cardiovascular disease mechanisms.
Biomarker Discovery: Sulfatides have emerged as potential biomarkers for various diseases due to their differential expression patterns in pathological conditions. Sulfatides analysis facilitates biomarker discovery efforts by identifying disease-specific sulfatide signatures in biofluids and tissues, offering non-invasive diagnostic and prognostic tools for precision medicine.
Drug Development: Sulfatides are targets for therapeutic intervention in diseases such as cancer and inflammatory disorders. Sulfatides analysis aids in drug discovery and development by evaluating the efficacy of sulfatide-targeting agents and elucidating their mechanisms of action in preclinical models and clinical trials.
Reference
- Cheng, Hua, et al. "Selective desorption/ionization of sulfatides by MALDI-MS facilitated using 9-aminoacridine as matrix." Journal of lipid research 51.6 (2010): 1599-1609.




